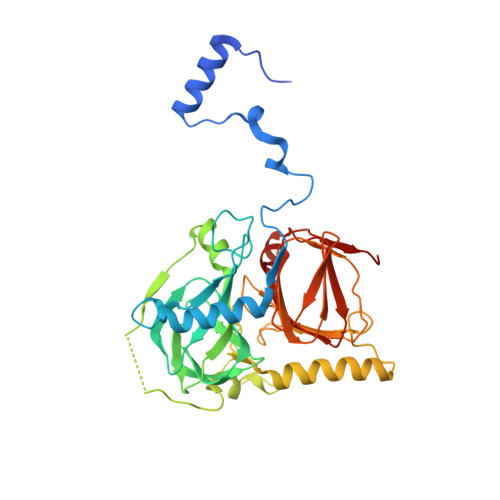
The salicylate 1,2-dioxygenase as a model for a conventional gentisate 1,2-dioxygenase: crystal structures of the G106A mutant and its adducts with gentisate and salicylate.
Ferraroni, M., Matera, I., Burger, S., Reichert, S., Steimer, L., Scozzafava, A., Stolz, A., Briganti, F.(2013) FEBS J 280: 1643-1652
- PubMed: 23384287 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12173
- Primary Citation Related Structures:
3NST, 3NVC, 3NW4 - PubMed Abstract:
The salicylate 1,2-dioxygenase (SDO) from the bacterium Pseudaminobacter salicylatoxidans BN12 is a versatile gentisate 1,2-dioxygenase (GDO) that converts both gentisate (2,5-dihydroxybenzoate) and various monohydroxylated substrates. Several variants of this enzyme were rationally designed based on the previously determined enzyme structure and sequence differences between the SDO and the 'conventional' GDO from Corynebacterium glutamicum. This was undertaken in order to define the structural elements that give the SDO its unique ability to dioxygenolytically cleave (substituted) salicylates. SDO variants M103L, G106A, G111A, R113G, S147R and F159Y were constructed and it was found that G106A oxidized only gentisate; 1-hydroxy-2-naphthoate and salicylate were not converted. This indicated that this enzyme variant behaves like previously known 'conventional' GDOs. Crystals of the G106A SDO variant and its complexes with salicylate and gentisate were obtained under anaerobic conditions, and the structures were solved and analyzed. The amino acid residue Gly106 is located inside the SDO active site cavity but does not directly interact with the substrates. Crystal structures of G106A SDO complexes with gentisate and salicylate showed a different binding mode for salicylate when compared with the wild-type enzyme. Thus, salicylate coordinated in the G106A variant with the catalytically active Fe(II) ion in an unusual and unproductive manner because of the inability of salicylate to displace a hydrogen bond that was formed between Trp104 and Asp174 in the G106A variant. It is proposed that this type of unproductive substrate binding might generally limit the substrate spectrum of 'conventional' GDOs. Structural data are available in the Protein Data Bank databases under the accession numbers 3NST, 3NWA, 3NVC.
- Dipartimento di Chimica Ugo Schiff, Università di Firenze, Sesto Fiorentino, Italy. marta.ferraroni@unifi.it
Organizational Affiliation: