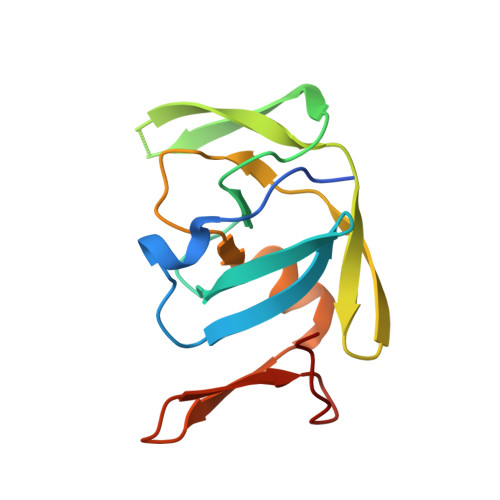
Crystal structure of XMRV protease differs from the structures of other retropepsins.
Li, M., Dimaio, F., Zhou, D., Gustchina, A., Lubkowski, J., Dauter, Z., Baker, D., Wlodawer, A.(2011) Nat Struct Mol Biol 18: 227-229
- PubMed: 21258323 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1964
- Primary Citation Related Structures:
3NR6 - PubMed Abstract:
Using energy and density guided Rosetta refinement to improve molecular replacement, we determined the crystal structure of the protease encoded by xenotropic murine leukemia virus-related virus (XMRV). Despite overall similarity of XMRV protease to other retropepsins, the topology of its dimer interface more closely resembles those of the monomeric, pepsin-like enzymes. Thus, XMRV protease may represent a distinct branch of the aspartic protease family.
- Protein Structure Section, Macromolecular Crystallography Laboratory, National Cancer Institute at Frederick, Frederick, Maryland, USA.
Organizational Affiliation: