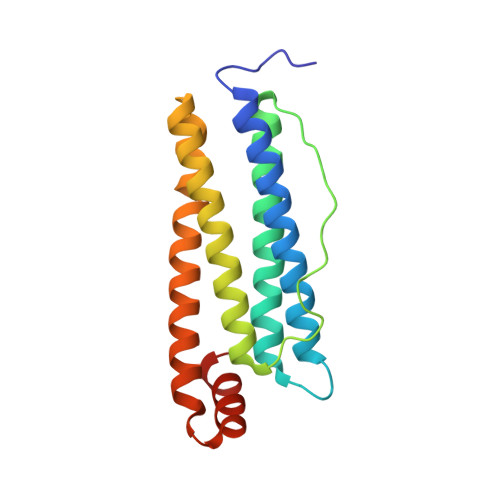
Definite coordination arrangement of organometallic palladium complexes accumulated on the designed interior surface of apo-ferritin.
Wang, Z., Takezawa, Y., Aoyagi, H., Abe, S., Hikage, T., Watanabe, Y., Kitagawa, S., Ueno, T.(2011) Chem Commun (Camb) 47: 170-172
- PubMed: 20730233 Search on PubMed
- DOI: https://doi.org/10.1039/c0cc02221g
- Primary Citation Related Structures:
3NOZ, 3NP0, 3NP2 - PubMed Abstract:
Apo-ferritin (apo-Fr) mutants are used as scaffolds to accommodate palladium (allyl) complexes. Various coordination arrangements of the Pd complexes are achieved by adjusting the positions of cysteine and histidine residues on the interior surface of the apo-Fr cage.
- Department of Chemistry, Graduate School of Science, Nagoya University, Furo-cho, Nagoya, 464-8602, Japan.
Organizational Affiliation: