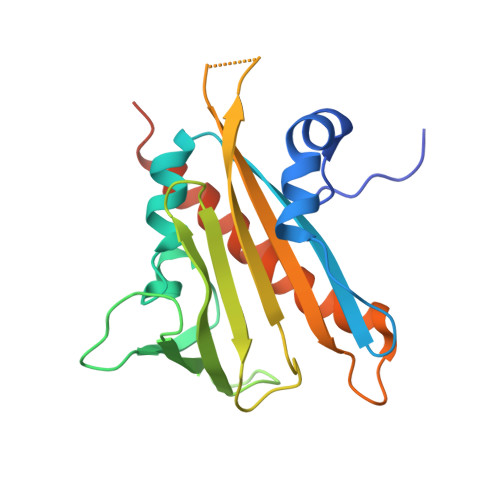
Functional mechanism of the ABA agonist pyrabactin
Hao, Q., Yin, P., Yan, C., Yuan, X., Li, W., Zhang, Z., Liu, L., Wang, J., Yan, N.(2010) J Biological Chem 285: 28946-28952
- PubMed: 20554531 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.149005
- Primary Citation Related Structures:
3NEF, 3NEG - PubMed Abstract:
Pyrabactin is a synthetic abscisic acid (ABA) agonist that selectively inhibits seed germination. The use of pyrabactin was pivotal in the identification of the PYR1/PYL/RCAR family (PYL) of proteins as the ABA receptor. Although they both act through PYL proteins, pyrabactin and ABA share no apparent chemical or structural similarity. It remains unclear how pyrabactin functions as an ABA agonist. Here, we report the crystal structure of pyrabactin in complex with PYL1 at 2.4 A resolution. Structural and biochemical analyses revealed that recognition of pyrabactin by the pocket residues precedes the closure of switch loop CL2. Structural comparison between pyrabactin- and ABA-bound PYL1 reveals a general principle in the arrangements of function groups of the two distinct ligands. The study provides a framework for the development of novel ABA agonists that may have applicable potentials in agriculture.
- State Key Laboratory of Biomembrane and Membrane Biotechnology, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: