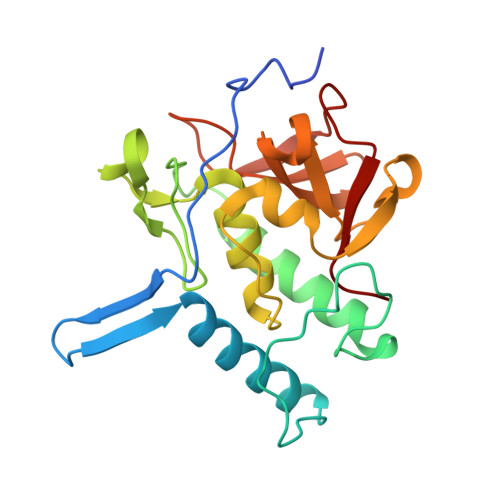
Structure and Functional Regulation of RipA, a Mycobacterial Enzyme Essential for Daughter Cell Separation.
Ruggiero, A., Marasco, D., Squeglia, F., Soldini, S., Pedone, E., Pedone, C., Berisio, R.(2010) Structure 18: 1184-1190
- PubMed: 20826344 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2010.06.007
- Primary Citation Related Structures:
3NE0 - PubMed Abstract:
Cell separation depends on cell-wall hydrolases that cleave the peptidoglycan layer connecting daughter cells. In Mycobacterium tuberculosis, this process is governed by the predicted endopeptidase RipA. In the absence of this enzyme, the bacterium is unable to divide and exhibits an abnormal phenotype. We here report the crystal structure of a relevant portion of RipA, containing its catalytic-domain and an extra-domain of hitherto unknown function. The structure clearly demonstrates that RipA is produced as a zymogen, which needs to be activated to achieve cell-division. Bacterial cell-wall degradation assays and proteolysis experiments strongly suggest that activation occurs via proteolytic processing of a fully solvent exposed loop identified in the crystal structure. Indeed, proteolytic cleavage at this loop produces an activated form, consisting of the sole catalytic domain. Our work provides the first evidence of self-inhibition in cell-disconnecting enzymes and opens a field for the design of novel antitubercular therapeutics.
- Institute of Biostructures and Bioimaging, CNR, Via Mezzocannone 16, I-80134, Naples, Italy.
Organizational Affiliation: