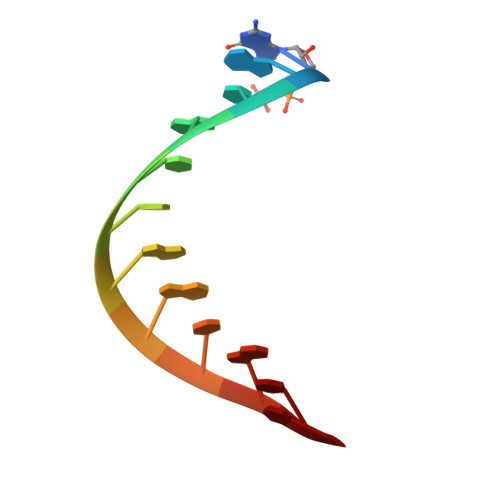
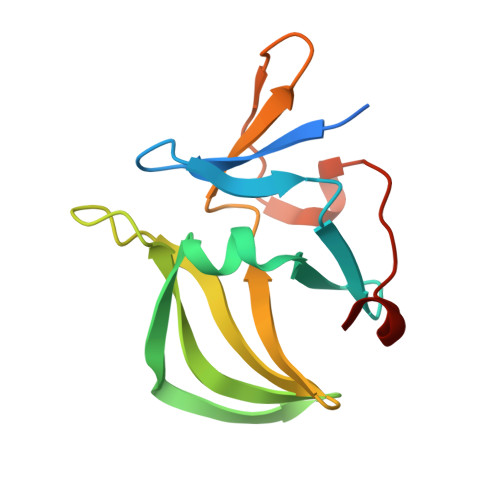
Structural and functional insights into 5'-ppp RNA pattern recognition by the innate immune receptor RIG-I.
Wang, Y., Ludwig, J., Schuberth, C., Goldeck, M., Schlee, M., Li, H., Juranek, S., Sheng, G., Micura, R., Tuschl, T., Hartmann, G., Patel, D.J.(2010) Nat Struct Mol Biol 17: 781-787
- PubMed: 20581823 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1863
- Primary Citation Related Structures:
3NCU - PubMed Abstract:
RIG-I is a cytosolic helicase that senses 5'-ppp RNA contained in negative-strand RNA viruses and triggers innate antiviral immune responses. Calorimetric binding studies established that the RIG-I C-terminal regulatory domain (CTD) binds to blunt-end double-stranded 5'-ppp RNA a factor of 17 more tightly than to its single-stranded counterpart. Here we report on the crystal structure of RIG-I CTD bound to both blunt ends of a self-complementary 5'-ppp dsRNA 12-mer, with interactions involving 5'-pp clearly visible in the complex. The structure, supported by mutation studies, defines how a lysine-rich basic cleft within the RIG-I CTD sequesters the observable 5'-pp of the bound RNA, with a stacked phenylalanine capping the terminal base pair. Key intermolecular interactions observed in the crystalline state are retained in the complex of 5'-ppp dsRNA 24-mer and full-length RIG-I under in vivo conditions, as evaluated from the impact of binding pocket RIG-I mutations and 2'-OCH(3) RNA modifications on the interferon response.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York, USA.
Organizational Affiliation: