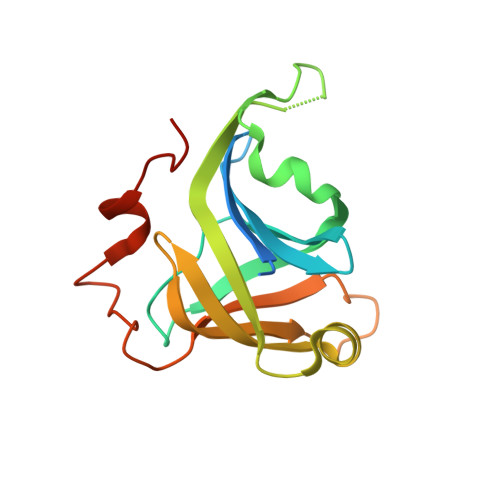
Structure of bacterial LigD 3'-phosphoesterase unveils a DNA repair superfamily
Nair, P.A., Smith, P., Shuman, S.(2010) Proc Natl Acad Sci U S A 107: 12822-12827
- PubMed: 20616014 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1005830107
- Primary Citation Related Structures:
3N9B, 3N9D - PubMed Abstract:
The DNA ligase D (LigD) 3'-phosphoesterase (PE) module is a conserved component of the bacterial nonhomologous end-joining (NHEJ) apparatus that performs 3' end-healing reactions at DNA double-strand breaks. Here we report the 1.9 A crystal structure of Pseudomonas aeruginosa PE, which reveals that PE exemplifies a unique class of DNA repair enzyme. PE has a distinctive fold in which an eight stranded beta barrel with a hydrophobic interior supports a crescent-shaped hydrophilic active site on its outer surface. Six essential side chains coordinate manganese and a sulfate mimetic of the scissile phosphate. The PE active site and mechanism are unique vis à vis other end-healing enzymes. We find PE homologs in archaeal and eukaryal proteomes, signifying that PEs comprise a DNA repair superfamily.
- Molecular Biology Program, Sloan-Kettering Institute, New York, NY 10065, USA.
Organizational Affiliation: