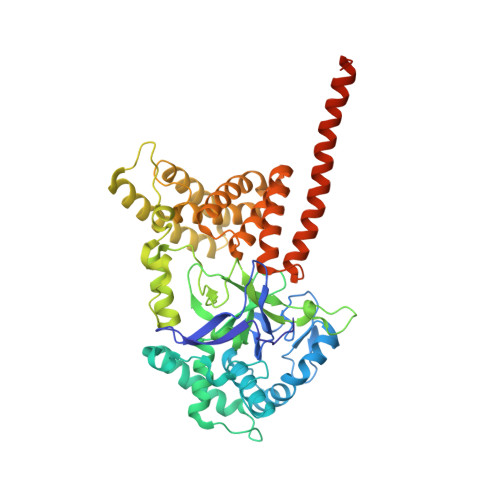
Crystal Structure of Cardiac-specific Histone Methyltransferase SmyD1 Reveals Unusual Active Site Architecture.
Sirinupong, N., Brunzelle, J., Ye, J., Pirzada, A., Nico, L., Yang, Z.(2010) J Biological Chem 285: 40635-40644
- PubMed: 20943667 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.168187
- Primary Citation Related Structures:
3N71 - PubMed Abstract:
SmyD1 is a cardiac- and muscle-specific histone methyltransferase that methylates histone H3 at lysine 4 and regulates gene transcription in early heart development. The unique domain structure characterized by a "split" SET domain, a conserved MYND zinc finger, and a novel C-terminal domain (CTD) distinguishes SmyD1 from other SET domain containing methyltransferases. Here we report the crystal structure of full-length SmyD1 in complex with the cofactor analog sinefungin at 2.3 Å. The structure reveals that SmyD1 folds into a wrench-shaped structure with two thick "grips" separated by a large, deep concave opening. Importantly, our structural and functional analysis suggests that SmyD1 appears to be regulated by an autoinhibition mechanism, and that unusually spacious target lysine-access channel and the presence of the CTD domain both negatively contribute to the regulation of this cardiovascularly relevant methyltransferase. Furthermore, our structure also provides a structural basis for the interaction between SmyD1 and cardiac transcription factor skNAC, and suggests that the MYND domain may primarily serve as a protein interaction module and cooperate SmyD1 with skNAC to regulate cardiomyocyte growth and maturation. Overall, our data provide novel insights into the mechanism of SmyD1 regulation, which would be helpful in further understanding the role of this protein in heart development and cardiovascular diseases.
- Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, Michigan 48201, USA.
Organizational Affiliation: