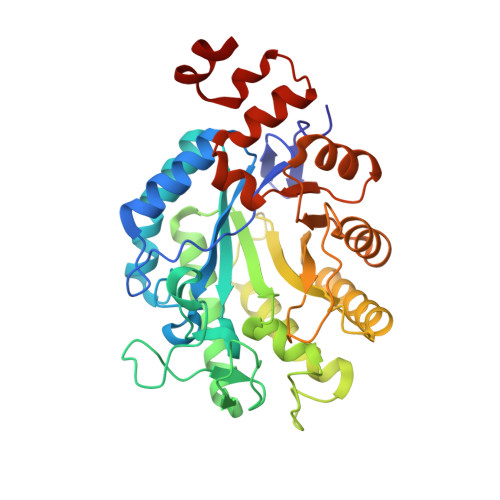
Determinants of substrate binding and protonation in the flavoenzyme xenobiotic reductase A
Spiegelhauer, O., Werther, T., Mende, S., Knauer, S.H., Dobbek, H.(2010) J Mol Biology 403: 286-298
- PubMed: 20826164 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.08.047
- Primary Citation Related Structures:
3N14, 3N19 - PubMed Abstract:
Xenobiotic reductase A (XenA) from Pseudomonas putida 86 catalyzes the NAD(P)H-dependent reduction of various α,β-unsaturated carbonyl compounds and is a member of the old yellow enzyme family. The reaction of XenA follows a ping-pong mechanism, implying that its active site has to accommodate and correctly position the various substrates to be oxidized (NADH/NADPH) and to be reduced (different α,β-unsaturated carbonyl compounds) to enable formal hydride transfers between the substrate and the isoalloxazine ring. The active site of XenA is lined by two tyrosine (Tyr27, Tyr183) and two tryptophan (Trp302, Trp358) residues, which were proposed to contribute to substrate binding. We analyzed the individual contributions of the four residues, using site-directed mutagenesis, steady-state and transient kinetics, redox potentiometry and crystal structure analysis. The Y183F substitution decreases the affinity of XenA for NADPH and reduces the rate of the oxidative half-reaction by two to three orders of magnitude, the latter being in agreement with its function as a proton donor in the oxidative half-reaction. Upon reduction of the flavin, Trp302 swings into the active site of XenA (in-conformation) and decreases the extent of the substrate-binding pocket. Its exchange against alanine induces substrate inhibition at elevated NADPH concentrations, indicating that the in-conformation of Trp302 helps to disfavor the nonproductive NADPH binding in the reduced state of XenA. Our analysis shows that while the principal catalytic mechanism of XenA, for example, type of proton donor, is analogous to that of other members of the old yellow enzyme family, its strategy to correctly position and accommodate different substrates is unprecedented.
- AG Bioanorganische Chemie, Universität Bayreuth, Universitätsstrasse 30, 95447 Bayreuth, Germany.
Organizational Affiliation: