Two Independent Histidines, One in Human Prolactin and One in Its Receptor, Are Critical for pH-dependent Receptor Recognition and Activation.
Kulkarni, M.V., Tettamanzi, M.C., Murphy, J.W., Keeler, C., Myszka, D.G., Chayen, N.E., Lolis, E.J., Hodsdon, M.E.(2010) J Biological Chem 285: 38524-38533
- PubMed: 20889499 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.172072
- Primary Citation Related Structures:
3MZG, 3N06, 3N0P, 3NCB, 3NCC, 3NCE, 3NCF - PubMed Abstract:
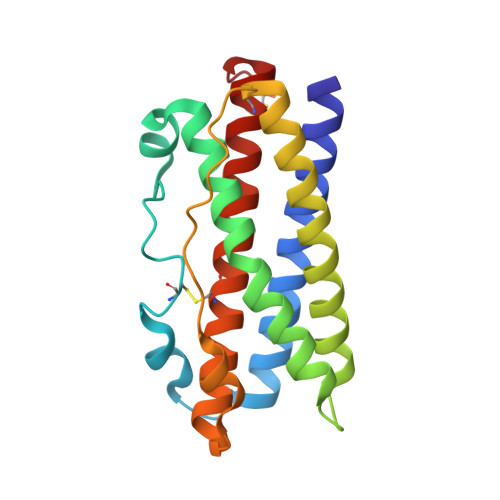
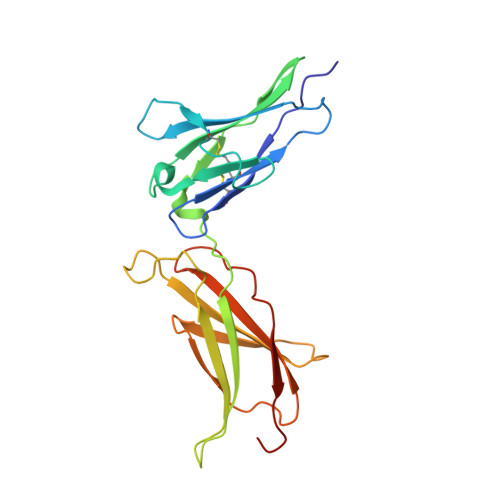
Human prolactin (hPRL), a member of the family of hematopoietic cytokines, functions as both an endocrine hormone and autocrine/paracrine growth factor. We have previously demonstrated that recognition of the hPRL·receptor depends strongly on solution acidity over the physiologic range from pH 6 to pH 8. The hPRL·receptor binding interface contains four histidines whose protonation is hypothesized to regulate pH-dependent receptor recognition. Here, we systematically dissect its molecular origin by characterizing the consequences of His to Ala mutations on pH-dependent receptor binding kinetics, site-specific histidine protonation, and high resolution structures of the intermolecular interface. Thermodynamic modeling of the pH dependence to receptor binding affinity reveals large changes in site-specific protonation constants for a majority of interface histidines upon complexation. Removal of individual His imidazoles reduces these perturbations in protonation constants, which is most likely explained by the introduction of solvent-filled, buried cavities in the crystallographic structures without inducing significant conformational rearrangements.
- Department of Laboratory Medicine, Yale School of Medicine, New Haven, Connecticut 06520, USA.
Organizational Affiliation: