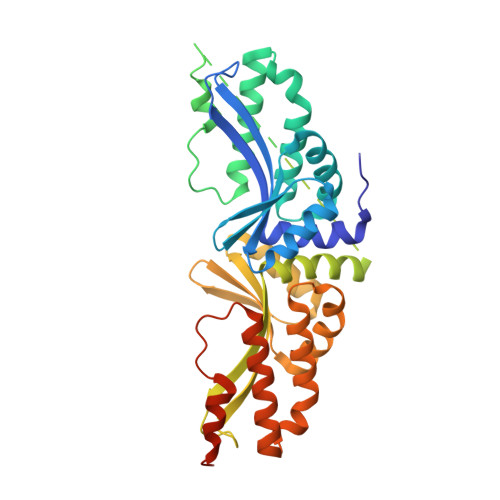
Molecular basis of engineered meganuclease targeting of the endogenous human RAG1 locus.
Munoz, I.G., Prieto, J., Subramanian, S., Coloma, J., Redondo, P., Villate, M., Merino, N., Marenchino, M., D'Abramo, M., Gervasio, F.L., Grizot, S., Daboussi, F., Smith, J., Chion-Sotinel, I., Paques, F., Duchateau, P., Alibes, A., Stricher, F., Serrano, L., Blanco, F.J., Montoya, G.(2011) Nucleic Acids Res 39: 729-743
- PubMed: 20846960 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq801
- Primary Citation Related Structures:
2XE0, 3MX9, 3MXA, 3MXB - PubMed Abstract:
Homing endonucleases recognize long target DNA sequences generating an accurate double-strand break that promotes gene targeting through homologous recombination. We have modified the homodimeric I-CreI endonuclease through protein engineering to target a specific DNA sequence within the human RAG1 gene. Mutations in RAG1 produce severe combined immunodeficiency (SCID), a monogenic disease leading to defective immune response in the individuals, leaving them vulnerable to infectious diseases. The structures of two engineered heterodimeric variants and one single-chain variant of I-CreI, in complex with a 24-bp oligonucleotide of the human RAG1 gene sequence, show how the DNA binding is achieved through interactions in the major groove. In addition, the introduction of the G19S mutation in the neighborhood of the catalytic site lowers the reaction energy barrier for DNA cleavage without compromising DNA recognition. Gene-targeting experiments in human cell lines show that the designed single-chain molecule preserves its in vivo activity with higher specificity, further enhanced by the G19S mutation. This is the first time that an engineered meganuclease variant targets the human RAG1 locus by stimulating homologous recombination in human cell lines up to 265 bp away from the cleavage site. Our analysis illustrates the key features for à la carte procedure in protein-DNA recognition design, opening new possibilities for SCID patients whose illness can be treated ex vivo.
- Structural Biology and Biocomputing Programme, Spanish National Cancer Research Centre (CNIO), Madrid, Spain.
Organizational Affiliation: