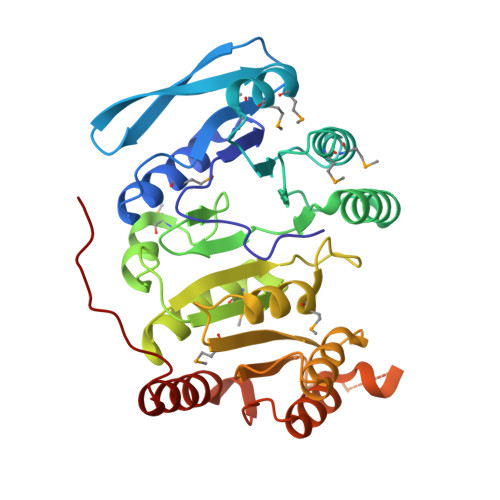
Identification of the citrate-binding site of human ATP-citrate lyase using X-ray crystallography.
Sun, T., Hayakawa, K., Bateman, K.S., Fraser, M.E.(2010) J Biological Chem 285: 27418-27428
- PubMed: 20558738 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.078667
- Primary Citation Related Structures:
3MWD, 3MWE - PubMed Abstract:
ATP-citrate lyase (ACLY) catalyzes the conversion of citrate and CoA into acetyl-CoA and oxaloacetate, coupled with the hydrolysis of ATP. In humans, ACLY is the cytoplasmic enzyme linking energy metabolism from carbohydrates to the production of fatty acids. In situ proteolysis of full-length human ACLY gave crystals of a truncated form, revealing the conformations of residues 2-425, 487-750, and 767-820 of the 1101-amino acid protein. Residues 2-425 form three domains homologous to the beta-subunit of succinyl-CoA synthetase (SCS), while residues 487-820 form two domains homologous to the alpha-subunit of SCS. The crystals were grown in the presence of tartrate or the substrate, citrate, and the structure revealed the citrate-binding site. A loop formed by residues 343-348 interacts via specific hydrogen bonds with the hydroxyl and carboxyl groups on the prochiral center of citrate. Arg-379 forms a salt bridge with the pro-R carboxylate of citrate. The pro-S carboxylate is free to react, providing insight into the stereospecificity of ACLY. Because this is the first structure of any member of the acyl-CoA synthetase (NDP-forming) superfamily in complex with its organic acid substrate, locating the citrate-binding site is significant for understanding the catalytic mechanism of each member, including the prototype SCS. Comparison of the CoA-binding site of SCSs with the similar structure in ACLY showed that ACLY possesses a different CoA-binding site. Comparisons of the nucleotide-binding site of SCSs with the similar structure in ACLY indicates that this is the ATP-binding site of ACLY.
- Department of Biological Sciences, University of Calgary, Calgary, Alberta T2N 1N4, Canada.
Organizational Affiliation: