Crystal structure determination of Japanese quail (Coturnix coturnix japonica) hemoglobin at 2.76 Angstrom resolution
Thenmozhi, M., Sathya Moorthy, P., Balasubramanian, M., Ponnuswamy, M.N.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hemoglobin subunit alpha-A | 141 | Coturnix japonica | Mutation(s): 0 | 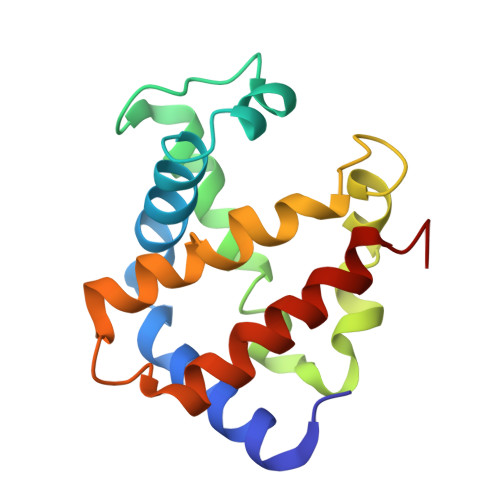 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P24589 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hemoglobin subunit beta | 146 | Coturnix japonica | Mutation(s): 0 | 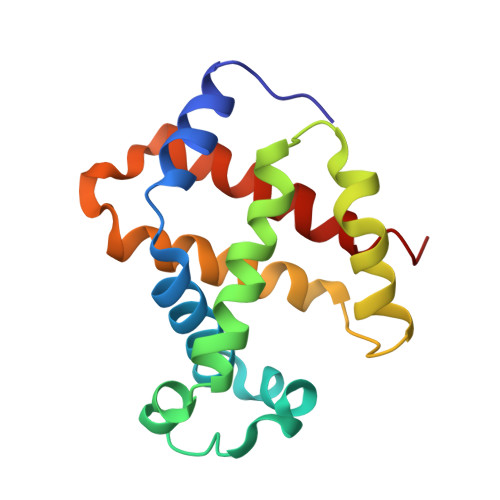 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P30893 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEM Download:Ideal Coordinates CCD File | E [auth A], G [auth B], I [auth C], K [auth D] | PROTOPORPHYRIN IX CONTAINING FE C34 H32 Fe N4 O4 KABFMIBPWCXCRK-RGGAHWMASA-L |  | ||
| OXY Download:Ideal Coordinates CCD File | F [auth A], H [auth B], J [auth C] | OXYGEN MOLECULE O2 MYMOFIZGZYHOMD-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 62.09 | α = 90 |
| b = 97.434 | β = 123.1 |
| c = 61.705 | γ = 90 |
| Software Name | Purpose |
|---|---|
| MAR345dtb | data collection |
| PHASER | phasing |
| REFMAC | refinement |
| AUTOMAR | data reduction |
| SCALEPACK | data scaling |