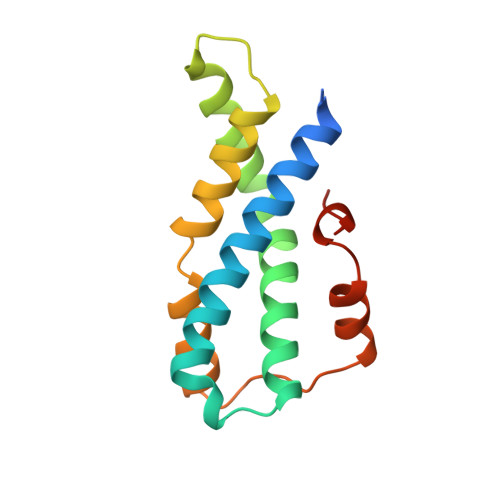
The high resolution crystal structure of green abalone sperm lysin: implications for species-specific binding of the egg receptor.
Kresge, N., Vacquier, V.D., Stout, C.D.(2000) J Mol Biology 296: 1225-1234
- PubMed: 10698629 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3533
- Primary Citation Related Structures:
3LYN - PubMed Abstract:
Abalone sperm lysin is a 16 kDa acrosomal protein used by sperm to create a hole in the egg vitelline envelope. Lysins from seven California abalone exhibit species-specificity in binding to their egg receptor, and range in sequence identity from 63 % to 90 %. The crystal structure of the sperm lysin dimer from Haliotis fulgens (green abalone) has been determined to 1.71 A by multiple isomorphous replacement. Comparisons with the structure of the lysin dimer from Haliotis rufescens (red abalone) reveal a similar overall fold and conservation of features contributing to lysin's amphipathic character. The two structures do, however, exhibit differences in surface residues and electrostatics. A large clustering of non-conserved surface residues around the waist and clefts of the dimer, and differences in charged residues around these regions, indicate areas of the molecule which may be involved in species-specific egg recognition.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037-1093, USA.
Organizational Affiliation: