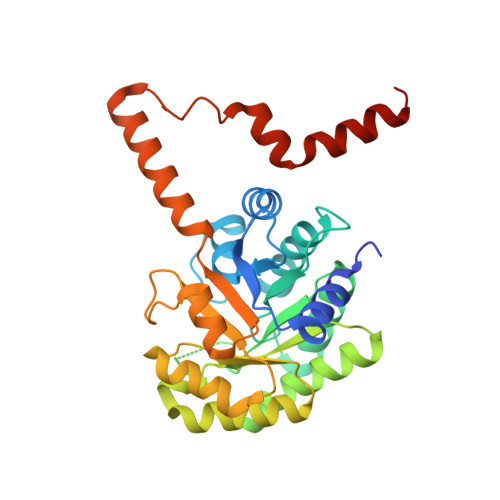
Structure of oxalacetate acetylhydrolase, a virulence factor of the chestnut blight fungus.
Chen, C., Sun, Q., Narayanan, B., Nuss, D.L., Herzberg, O.(2010) J Biological Chem 285: 26685-26696
- PubMed: 20558740 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.117804
- Primary Citation Related Structures:
3LYE, 3M0J, 3M0K - PubMed Abstract:
Oxalacetate acetylhydrolase (OAH), a member of the phosphoenolpyruvate mutase/isocitrate lyase superfamily, catalyzes the hydrolysis of oxalacetate to oxalic acid and acetate. This study shows that knock-out of the oah gene in Cryphonectria parasitica, the chestnut blight fungus, reduces the ability of the fungus to form cankers on chestnut trees, suggesting that OAH plays a key role in virulence. OAH was produced in Escherichia coli and purified, and its catalytic rates were determined. Oxalacetate is the main OAH substrate, but the enzyme also acts as a lyase of (2R,3S)-dimethyl malate with approximately 1000-fold lower efficacy. The crystal structure of OAH was determined alone, in complex with a mechanism-based inhibitor, 3,3-difluorooxalacetate (DFOA), and in complex with the reaction product, oxalate, to a resolution limit of 1.30, 1.55, and 1.65 A, respectively. OAH assembles into a dimer of dimers with each subunit exhibiting an (alpha/beta)(8) barrel fold and each pair swapping the 8th alpha-helix. An active site "gating loop" exhibits conformational disorder in the ligand-free structure. To obtain the structures of the OAH.ligand complexes, the ligand-free OAH crystals were soaked briefly with DFOA or oxalacetate. DFOA binding leads to ordering of the gating loop in a conformation that sequesters the ligand from the solvent. DFOA binds in a gem-diol form analogous to the oxalacetate intermediate/transition state. Oxalate binds in a planar conformation, but the gating loop is largely disordered. Comparison between the OAH structure and that of the closely related enzyme, 2,3-dimethylmalate lyase, suggests potential determinants of substrate preference.
- W. M. Keck Laboratory for Structural Biology, Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, Rockville, Maryland 20850, USA.
Organizational Affiliation: