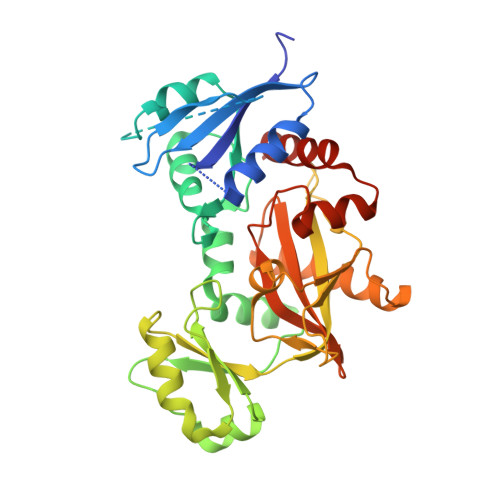
Structure of the Mycobacterium tuberculosis D-Alanine:D-Alanine Ligase, a Target of the Antituberculosis Drug D-Cycloserine.
Bruning, J.B., Murillo, A.C., Chacon, O., Barletta, R.G., Sacchettini, J.C.(2011) Antimicrob Agents Chemother 55: 291-301
- PubMed: 20956591 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AAC.00558-10
- Primary Citation Related Structures:
3LWB - PubMed Abstract:
D-alanine:D-alanine ligase (EC 6.3.2.4; Ddl) catalyzes the ATP-driven ligation of two D-alanine (D-Ala) molecules to form the D-alanyl:D-alanine dipeptide. This molecule is a key building block in peptidoglycan biosynthesis, making Ddl an attractive target for drug development. D-Cycloserine (DCS), an analog of D-Ala and a prototype Ddl inhibitor, has shown promise for the treatment of tuberculosis. Here, we report the crystal structure of Mycobacterium tuberculosis Ddl at a resolution of 2.1 Å. This structure indicates that Ddl is a dimer and consists of three discrete domains; the ligand binding cavity is at the intersection of all three domains and conjoined by several loop regions. The M. tuberculosis apo Ddl structure shows a novel conformation that has not yet been observed in Ddl enzymes from other species. The nucleotide and D-alanine binding pockets are flexible, requiring significant structural rearrangement of the bordering regions for entry and binding of both ATP and D-Ala molecules. Solution affinity and kinetic studies showed that DCS interacts with Ddl in a manner similar to that observed for D-Ala. Each ligand binds to two binding sites that have significant differences in affinity, with the first binding site exhibiting high affinity. DCS inhibits the enzyme, with a 50% inhibitory concentration (IC(50)) of 0.37 mM under standard assay conditions, implicating a preferential and weak inhibition at the second, lower-affinity binding site. Moreover, DCS binding is tighter at higher ATP concentrations. The crystal structure illustrates potential drugable sites that may result in the development of more-effective Ddl inhibitors.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77843-2128, USA.
Organizational Affiliation: