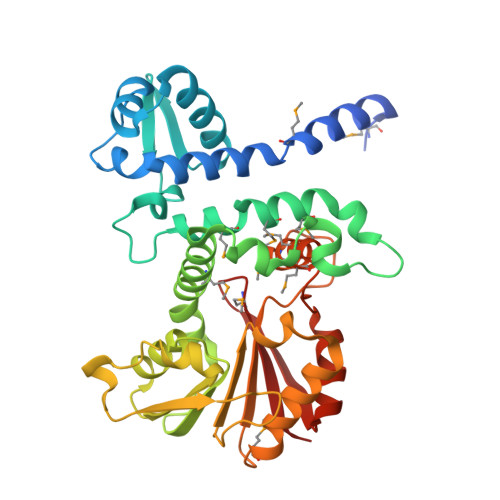
Structural characterization of CalO1: a putative orsellinic acid methyltransferase in the calicheamicin-biosynthetic pathway.
Chang, A., Singh, S., Bingman, C.A., Thorson, J.S., Phillips, G.N.(2011) Acta Crystallogr D Biol Crystallogr 67: 197-203
- PubMed: 21358050 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S090744491100360X
- Primary Citation Related Structures:
3LST - PubMed Abstract:
The X-ray structure determination at 2.4 Å resolution of the putative orsellinic acid C3 O-methyltransferase (CalO1) involved in calicheamicin biosynthesis is reported. Comparison of CalO1 with a homology model of the functionally related calicheamicin orsellinic acid C2 O-methyltransferase (CalO6) implicates several residues that are likely to contribute to the regiospecificity of alkylation. Consistent with the proposed requirement of an acyl-carrier-protein-bound substrate, this structural study also reveals structural determinants within CalO1 that are anticipated to accommodate an association with an acyl carrier protein.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, Wisconsin 53706, USA.
Organizational Affiliation: