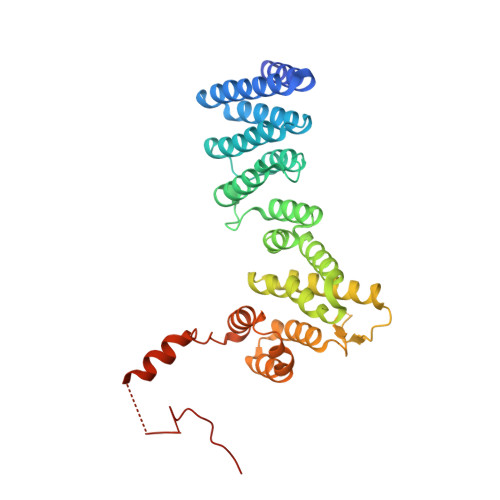
The structure of Get4 reveals an alpha-solenoid fold adapted for multiple interactions in tail-anchored protein biogenesis.
Bozkurt, G., Wild, K., Amlacher, S., Hurt, E., Dobberstein, B., Sinning, I.(2010) FEBS Lett 584: 1509-1514
- PubMed: 20206626 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2010.02.070
- Primary Citation Related Structures:
3LPZ - PubMed Abstract:
Tail-anchored proteins play important roles in protein translocation, membrane fusion and apoptosis. They are targeted to the endoplasmic reticulum membrane via the guided-entry of tail-anchored proteins (Get) pathway. We present the 2A crystal structure of Get4 which participates in early steps of the Get pathway. The structure shows an alpha-solenoid fold with particular deviations from the regular pairwise arrangement of alpha-helices. A conserved hydrophobic groove accommodates the flexible C-terminal region in trans. The structural organization of the Get4 helical hairpin motifs provides a scaffold for protein-protein interactions in the Get pathway.
- Heidelberg University Biochemistry Center (BZH), Heidelberg, Germany.
Organizational Affiliation: