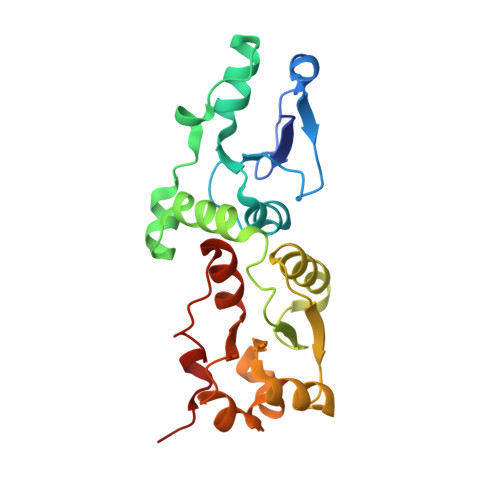
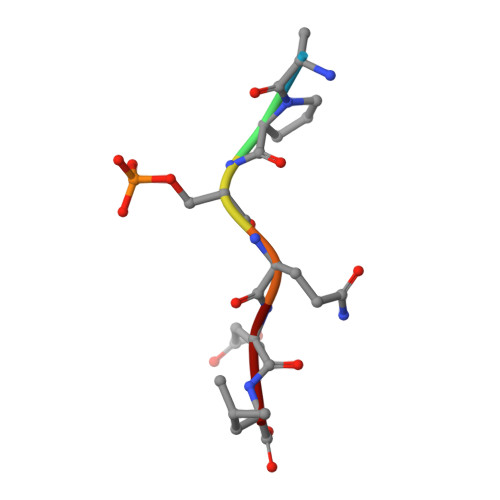
gammaH2A binds Brc1 to maintain genome integrity during S-phase.
Williams, J.S., Williams, R.S., Dovey, C.L., Guenther, G., Tainer, J.A., Russell, P.(2010) EMBO J 29: 1136-1148
- PubMed: 20094029 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2009.413
- Primary Citation Related Structures:
3L40, 3L41 - PubMed Abstract:
ATM(Tel1) and ATR(Rad3) checkpoint kinases phosphorylate the C-terminus of histone H2AX (H2A in yeasts) in chromatin flanking DNA damage, establishing a recruitment platform for checkpoint and repair proteins. Phospho-H2A/X (gammaH2A/X)-binding proteins at double-strand breaks (DSBs) have been characterized, but those required for replication stress responses are unknown. Here, we present genetic, biochemical, small angle X-ray scattering (SAXS), and X-ray structural studies of the Schizosaccharomyces pombe Brc1, a 6-BRCT-domain protein that is structurally related to Saccharomyces cerevisiae Rtt107 and mammalian PTIP. Brc1 binds gammaH2A to form spontaneous and DNA damage-induced nuclear foci. Spontaneous Brc1 foci colocalize with ribosomal DNA repeats, a region prone to fork pausing and genomic instability, whereas DNA damage-induced Brc1 foci colocalize with DSB response factors. gammaH2A binding is critical for Brc1 function. The 1.45 A resolution crystal structure of Brc1-gammaH2A complex shows how variable BRCT insertion loops sculpt tandem-BRCT phosphoprotein-binding pockets to facilitate unique phosphoprotein-interaction specificities, and unveils an acidic DNA-mimicking Brc1 surface. From these results, Brc1 docking to gammaH2A emerges as a critical chromatin-specific response to replication-associated DNA damage.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA, USA.
Organizational Affiliation: