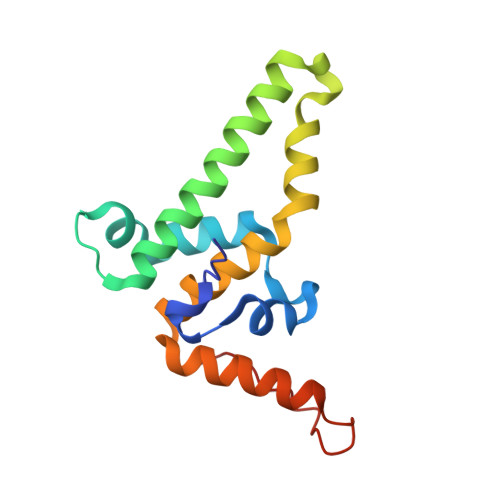
Conformational changes in the hepatitis B virus core protein are consistent with a role for allostery in virus assembly
Packianathan, C., Katen, S.P., Dann, C.E., Zlotnick, A.(2010) J Virol 84: 1607-1615
- PubMed: 19939922 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02033-09
- Primary Citation Related Structures:
3KXS - PubMed Abstract:
In infected cells, virus components must be organized at the right place and time to ensure assembly of infectious virions. From a different perspective, assembly must be prevented until all components are available. Hypothetically, this can be achieved by allosterically controlling assembly. Consistent with this hypothesis, here we show that the structure of the hepatitis B virus (HBV) core protein dimer, which can spontaneously self-assemble, is incompatible with capsid assembly. Systematic differences between core protein dimer and capsid conformations demonstrate linkage between the intradimer interface and interdimer contact surface. These structures also provide explanations for the capsid-dimer selectivity of some antibodies and the activities of assembly effectors. Solution studies suggest that the assembly-inactive state is more accurately an ensemble of conformations. Simulations show that allostery supports controlled assembly and results in capsids that are resistant to dissociation. We propose that allostery, as demonstrated in HBV, is common to most self-assembling viruses.
- Department of Molecular and Cellular Biochemistry, Indiana University, 212 S. Hawthorne Dr., Bloomington, IN 47405-7003, USA.
Organizational Affiliation: