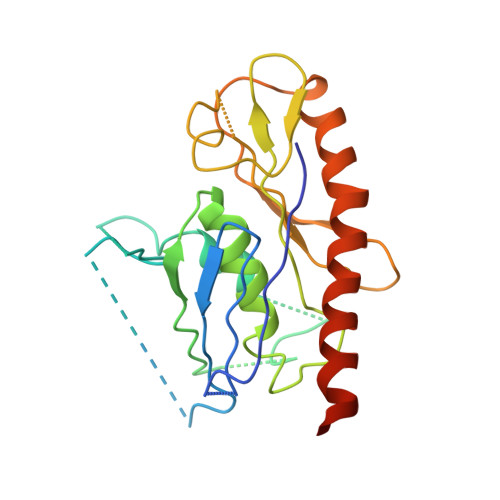
Structural and biochemical characterization of CRN-5 and Rrp46: an exosome component participating in apoptotic DNA degradation
Yang, C.-C., Wang, Y.-T., Hsiao, Y.-Y., Doudeva, L.G., Kuo, P.-H., Chow, S.Y., Yuan, H.S.(2010) RNA 16: 1748-1759
- PubMed: 20660080 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.2180810
- Primary Citation Related Structures:
3HKM, 3KRN - PubMed Abstract:
Rrp46 was first identified as a protein component of the eukaryotic exosome, a protein complex involved in 3' processing of RNA during RNA turnover and surveillance. The Rrp46 homolog, CRN-5, was subsequently characterized as a cell death-related nuclease, participating in DNA fragmentation during apoptosis in Caenorhabditis elegans. Here we report the crystal structures of CRN-5 and rice Rrp46 (oRrp46) at a resolution of 3.9 A and 2.0 A, respectively. We found that recombinant human Rrp46 (hRrp46), oRrp46, and CRN-5 are homodimers, and that endogenous hRrp46 and oRrp46 also form homodimers in a cellular environment, in addition to their association with a protein complex. Dimeric oRrp46 had both phosphorolytic RNase and hydrolytic DNase activities, whereas hRrp46 and CRN-5 bound to DNA without detectable nuclease activity. Site-directed mutagenesis in oRrp46 abolished either its DNase (E160Q) or RNase (K75E/Q76E) activities, confirming the critical importance of these residues in catalysis or substrate binding. Moreover, CRN-5 directly interacted with the apoptotic nuclease CRN-4 and enhanced the DNase activity of CRN-4, suggesting that CRN-5 cooperates with CRN-4 in apoptotic DNA degradation. Taken together all these results strongly suggest that Rrp46 forms a homodimer separately from exosome complexes and, depending on species, is either a structural or catalytic component of the machinery that cleaves DNA during apoptosis.
- Graduate Institute of Biochemistry and Molecular Biology, National Taiwan University, Taipei, 10617 Taiwan, Republic of China.
Organizational Affiliation: