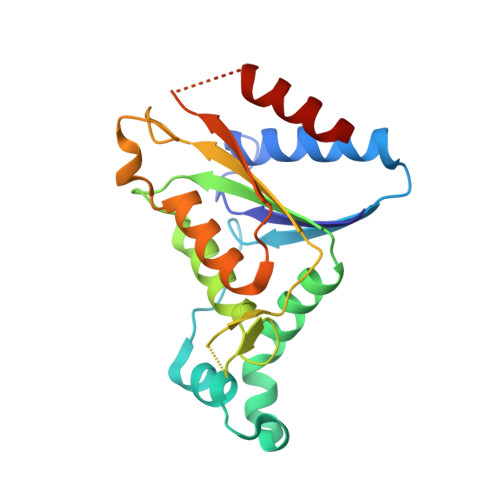
Role of tyrosine 131 in the active site of paAzoR1, an azoreductase with specificity for the inflammatory bowel disease prodrug balsalazide
Wang, C.-J., Laurieri, N., Abuhammad, A., Lowe, E., Westwood, I., Ryan, A., Sim, E.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 2-7
- PubMed: 20057057 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109044741
- Primary Citation Related Structures:
3KEG - PubMed Abstract:
Azoreductase 1 from Pseudomonas aeruginosa strain PAO1 (paAzoR1) catalyses the activation of the prodrug balsalazide and reduces the azo dye methyl red using reduced nicotinamide adenine dinucleotide cofactor as an electron donor. To investigate the mechanism of the enzyme, a Y131F mutation was introduced and the enzymic properties of the mutant were compared with those of the wild-type enzyme. The crystallographic structure of the mutant with methyl red bound was solved at 2.1 A resolution and compared with the wild-type structure. Tyr131 is important in the architecture of the active site but is not essential for enzymic activity.
- Department of Pharmacology, University of Oxford, Mansfield Road, Oxford OX1 3QT, England.
Organizational Affiliation: