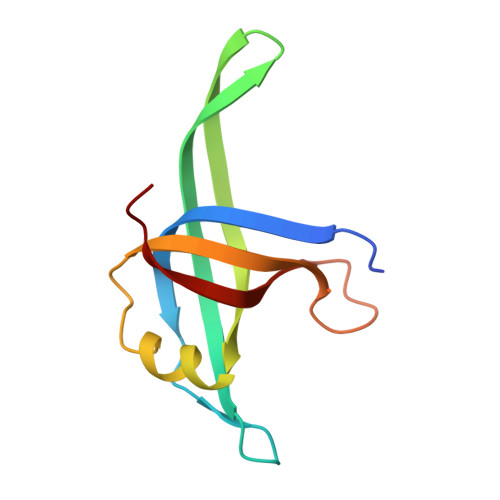
The crystal structure of Neisseria gonorrhoeae PriB reveals mechanistic differences among bacterial DNA replication restart pathways
Dong, J., George, N.P., Duckett, K.L., DeBeer, M.A., Lopper, M.E.(2010) Nucleic Acids Res 38: 499-509
- PubMed: 19906704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkp1031
- Primary Citation Related Structures:
3K8A - PubMed Abstract:
Reactivation of repaired DNA replication forks is essential for complete duplication of bacterial genomes. However, not all bacteria encode homologs of the well-studied Escherichia coli DNA replication restart primosome proteins, suggesting that there might be distinct mechanistic differences among DNA replication restart pathways in diverse bacteria. Since reactivation of repaired DNA replication forks requires coordinated DNA and protein binding by DNA replication restart primosome proteins, we determined the crystal structure of Neisseria gonorrhoeae PriB at 2.7 A resolution and investigated its ability to physically interact with DNA and PriA helicase. Comparison of the crystal structures of PriB from N. gonorrhoeae and E. coli reveals a well-conserved homodimeric structure consisting of two oligosaccharide/oligonucleotide-binding (OB) folds. In spite of their overall structural similarity, there is significant species variation in the type and distribution of surface amino acid residues. This correlates with striking differences in the affinity with which each PriB homolog binds single-stranded DNA and PriA helicase. These results provide evidence that mechanisms of DNA replication restart are not identical across diverse species and that these pathways have likely become specialized to meet the needs of individual organisms.
- Department of Chemistry, University of Dayton, 300 College Park, Dayton, OH 45469, USA.
Organizational Affiliation: