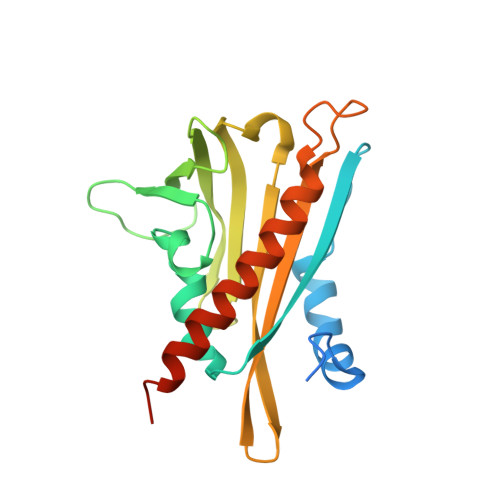
Structural mechanism of abscisic acid binding and signaling by dimeric PYR1.
Nishimura, N., Hitomi, K., Arvai, A.S., Rambo, R.P., Hitomi, C., Cutler, S.R., Schroeder, J.I., Getzoff, E.D.(2009) Science 326: 1373-1379
- PubMed: 19933100 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1181829
- Primary Citation Related Structures:
3K3K - PubMed Abstract:
The phytohormone abscisic acid (ABA) acts in seed dormancy, plant development, drought tolerance, and adaptive responses to environmental stresses. Structural mechanisms mediating ABA receptor recognition and signaling remain unknown but are essential for understanding and manipulating abiotic stress resistance. Here, we report structures of pyrabactin resistance 1 (PYR1), a prototypical PYR/PYR1-like (PYL)/regulatory component of ABA receptor (RCAR) protein that functions in early ABA signaling. The crystallographic structure reveals an alpha/beta helix-grip fold and homodimeric assembly, verified in vivo by coimmunoprecipitation. ABA binding within a large internal cavity switches structural motifs distinguishing ABA-free "open-lid" from ABA-bound "closed-lid" conformations. Small-angle x-ray scattering suggests that ABA signals by converting PYR1 to a more compact, symmetric closed-lid dimer. Site-directed PYR1 mutants designed to disrupt hormone binding lose ABA-triggered interactions with type 2C protein phosphatase partners in planta.
- Division of Biological Sciences, Cell and Developmental Biology Section, University of California at San Diego, La Jolla, CA 92093, USA.
Organizational Affiliation: