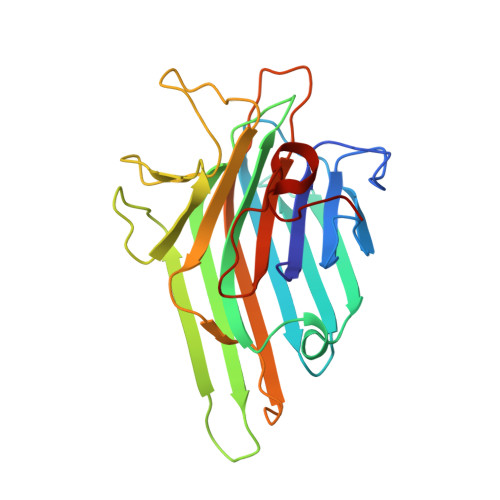
Structural analysis of ConBr reveals molecular correlation between the carbohydrate recognition domain and endothelial NO synthase activation.
Bezerra, E.H., Rocha, B.A., Nagano, C.S., Bezerra, G.A., Moura, T.R., Bezerra, M.J., Benevides, R.G., Sampaio, A.H., Assreuy, A.M., Delatorre, P., Cavada, B.S.(2011) Biochem Biophys Res Commun 408: 566-570
- PubMed: 21530490 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2011.04.061
- Primary Citation Related Structures:
3JU9 - PubMed Abstract:
Diocleinae lectins are highly homologous in their primary structure which features metal binding sites and a carbohydrate recognition domain (CRD). Differences in the biological activity of legume lectins have been widely investigated using hemagglutination inhibition assays, isothermal titration microcalorimetry and co-crystallization with mono- and oligosaccharides. Here we report a new lectin crystal structure (ConBr) extracted from seeds of Canavalia brasiliensis, predict dimannoside binding by docking, identify the α-aminobutyric acid (Abu) binding pocket and compare the CRD of ConBr to that of homologous lectins. Based on the hypothesis that the carbohydrate affinity of lectins depends on CRD configuration, the relationship between tridimensional structure and endothelial NO synthase activation was used to clarify differences in biological activity. Our study established a correlation between the position of CRD amino acid side chains and the stimulation of NO release from endothelium.
- Departamento de Bioquímica e Biologia Molecular, Universidade Federal do Ceará, Fortaleza, CE, Brazil.
Organizational Affiliation: