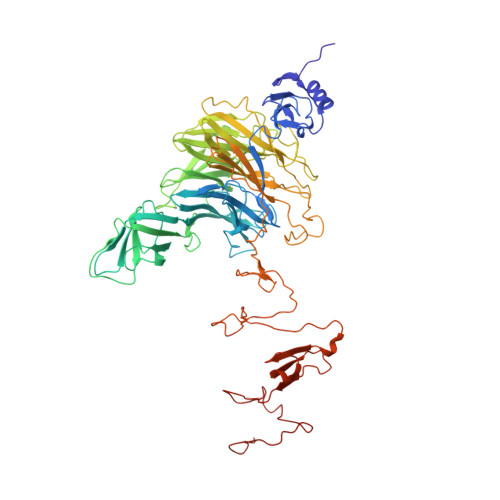
Structure analysis of endosialidase NF at 0.98 A resolution.
Schulz, E.C., Neumann, P., Gerardy-Schahn, R., Sheldrick, G.M., Ficner, R.(2010) Acta Crystallogr D Biol Crystallogr 66: 176-180
- PubMed: 20124697 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444909048720
- Primary Citation Related Structures:
3JU4 - PubMed Abstract:
Endosialidase NF (endoNF) is a bacteriophage-derived endosialidase that specifically degrades alpha-2,8-linked polysialic acid. The structure of a new crystal form of endoNF in complex with sialic acid has been refined at 0.98 A resolution. The 210 kDa homotrimeric multi-domain enzyme displays outstanding stability and resistance to SDS. Even at atomic resolution, only a minor fraction of side chains possess alternative conformations. However, multiple conformations of an active-site residue imply that it has an important catalytic function in the cleavage mechanism of polysialic acid.
- Abteilung für Molekulare Strukturbiologie, Institut für Mikrobiologie und Genetik, Georg-August-Universität Göttingen, Justus-von-Liebig-Weg 11, 37077 Göttingen, Germany.
Organizational Affiliation: