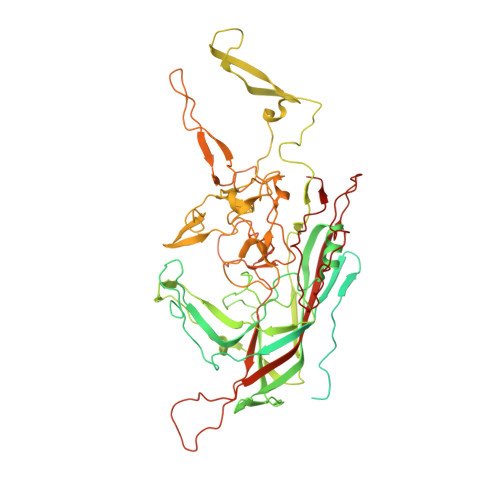
Structure of AAV-DJ, a retargeted gene therapy vector: cryo-electron microscopy at 4.5 A resolution.
Lerch, T.F., O'Donnell, J.K., Meyer, N.L., Xie, Q., Taylor, K.A., Stagg, S.M., Chapman, M.S.(2012) Structure 20: 1310-1320
- PubMed: 22727812 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.05.004
- Primary Citation Related Structures:
3J1Q - PubMed Abstract:
AAV-DJ, a leading candidate vector for liver gene therapy, was created through random homologous recombination followed by directed evolution, selecting for in vivo liver tropism and resistance to in vitro immune neutralization. Here, the 4.5 Å resolution cryo-EM structure is determined for the engineered AAV vector, revealing structural features that illuminate its phenotype. The heparan sulfate receptor-binding site is little changed from AAV-2, and heparin-binding affinity is similar. A loop that is antigenic in other serotypes has a unique conformation in AAV-DJ that would conflict with the binding of an AAV-2 neutralizing monoclonal antibody. This is consistent with increased resistance to neutralization by human polyclonal sera, raising the possibility that changed tropism may be a secondary effect of altered immune interactions. The reconstruction exemplifies analysis of fine structural changes and the potential of cryo-EM, in favorable cases, to characterize mutant or ligand-bound complexes.
- Department of Biochemistry and Molecular Biology, School of Medicine, Oregon Health and Science University, Portland, OR 97239, USA.
Organizational Affiliation: