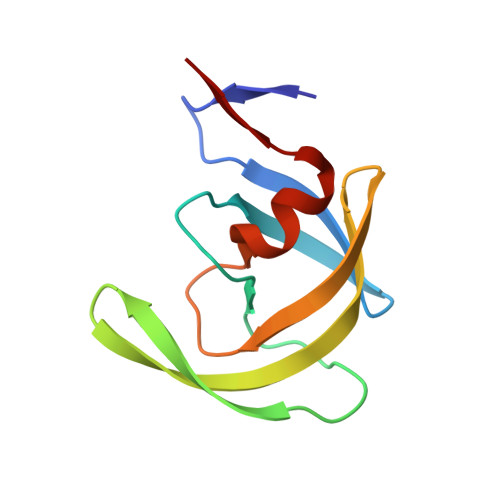
Structure of the unbound form of HIV-1 subtype A protease: comparison with unbound forms of proteases from other HIV subtypes.
Robbins, A.H., Coman, R.M., Bracho-Sanchez, E., Fernandez, M.A., Gilliland, C.T., Li, M., Agbandje-McKenna, M., Wlodawer, A., Dunn, B.M., McKenna, R.(2010) Acta Crystallogr D Biol Crystallogr 66: 233-242
- PubMed: 20179334 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444909054298
- Primary Citation Related Structures:
3IXO - PubMed Abstract:
The crystal structure of the unbound form of HIV-1 subtype A protease (PR) has been determined to 1.7 A resolution and refined as a homodimer in the hexagonal space group P6(1) to an R(cryst) of 20.5%. The structure is similar in overall shape and fold to the previously determined subtype B, C and F PRs. The major differences lie in the conformation of the flap region. The flaps in the crystal structures of the unbound subtype B and C PRs, which were crystallized in tetragonal space groups, are either semi-open or wide open. In the present structure of subtype A PR the flaps are found in the closed position, a conformation that would be more anticipated in the structure of HIV protease complexed with an inhibitor. The amino-acid differences between the subtypes and their respective crystal space groups are discussed in terms of the differences in the flap conformations.
- Department of Biochemistry and Molecular Biology, University of Florida, Gainesville, FL 32610, USA.
Organizational Affiliation: