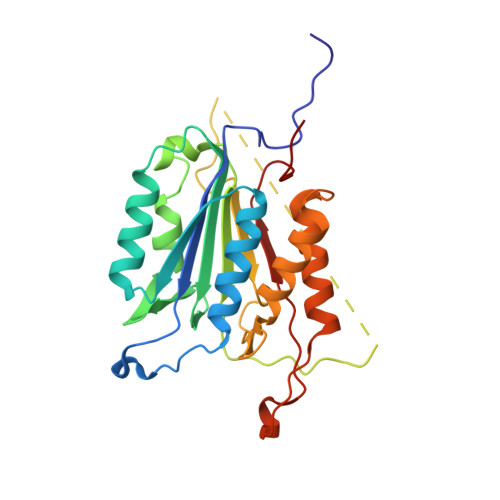
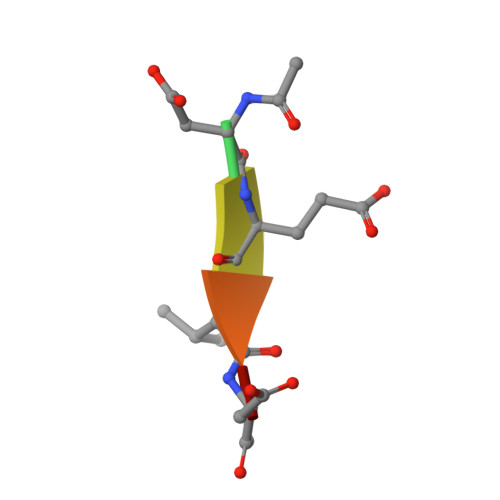
A constitutively active and uninhibitable caspase-3 zymogen efficiently induces apoptosis.
Walters, J., Pop, C., Scott, F.L., Drag, M., Swartz, P., Mattos, C., Salvesen, G.S., Clark, A.C.(2009) Biochem J 424: 335-345
- PubMed: 19788411 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20090825
- Primary Citation Related Structures:
3ITN - PubMed Abstract:
The caspase-3 zymogen has essentially zero activity until it is cleaved by initiator caspases during apoptosis. However, a mutation of V266E in the dimer interface activates the protease in the absence of chain cleavage. We show that low concentrations of the pseudo-activated procaspase-3 kill mammalian cells rapidly and, importantly, this protein is not cleaved nor is it inhibited efficiently by the endogenous regulator XIAP (X-linked inhibitor of apoptosis). The 1.63 A (1 A = 0.1 nm) structure of the variant demonstrates that the mutation is accommodated at the dimer interface to generate an enzyme with substantially the same activity and specificity as wild-type caspase-3. Structural modelling predicts that the interface mutation prevents the intersubunit linker from binding in the dimer interface, allowing the active sites to form in the procaspase in the absence of cleavage. The direct activation of procaspase-3 through a conformational switch rather than by chain cleavage may lead to novel therapeutic strategies for inducing cell death.
- Department of Molecular and Structural Biochemistry, North Carolina State University, Raleigh, NC 27695, USA.
Organizational Affiliation: