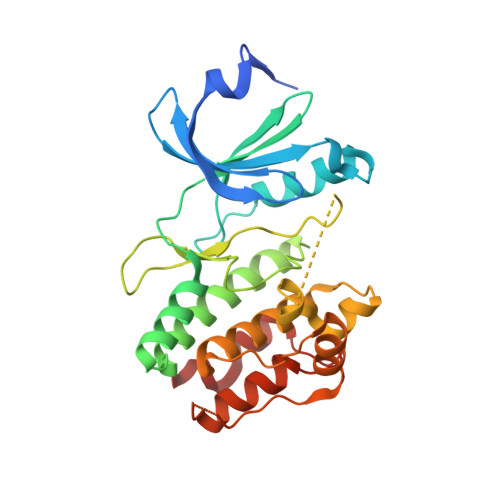
Crystal structure of CDPK kinase domain from toxoplasma Gondii, TGME49_018720
Wernimont, A.K., Artz, J.D., Senisterra, G., MacKenzie, F., Hutchinson, A., Kozieradzki, I., Cossar, D., Bochkarev, A., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Weigelt, J., Hui, R., Lin, Y.H.To be published.