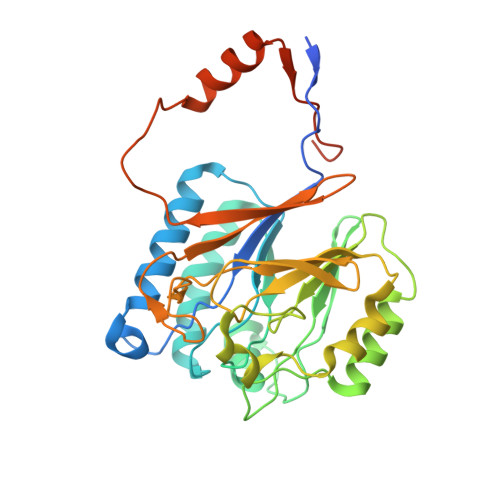
A mycobacterial cyclic AMP phosphodiesterase that moonlights as a modifier of cell wall permeability
Podobnik, M., Tyagi, R., Matange, N., Dermol, U., Gupta, A.K., Mattoo, R., Seshadri, K., Visweswariah, S.S.(2009) J Biological Chem 284: 32846-32857
- PubMed: 19801656 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.049635
- Primary Citation Related Structures:
3IB7, 3IB8 - PubMed Abstract:
Mycobacterium tuberculosis utilizes many mechanisms to establish itself within the macrophage, and bacterially derived cAMP is important in modulating the host cellular response. Although the genome of M. tuberculosis is endowed with a number of mammalian-like adenylyl cyclases, only a single cAMP phosphodiesterase has been identified that can decrease levels of cAMP produced by the bacterium. We present the crystal structure of the full-length and sole cAMP phosphodiesterase, Rv0805, found in M. tuberculosis, whose orthologs are present only in the genomes of slow growing and pathogenic mycobacteria. The dimeric core catalytic domain of Rv0805 adopts a metallophosphoesterase-fold, and the C-terminal region builds the active site and contributes to multiple substrate utilization. Localization of Rv0805 to the cell wall is dependent on its C terminus, and expression of either wild type or mutationally inactivated Rv0805 in M. smegmatis alters cell permeability to hydrophobic cytotoxic compounds. Rv0805 may therefore play a key role in the pathogenicity of mycobacteria, not only by hydrolyzing bacterial cAMP, but also by moonlighting as a protein that can alter cell wall functioning.
- Laboratory for Biosynthesis and Biotransformation, National Institute of Chemistry of Slovenia, Hajdrihova 19,1000 Ljubljana, Slovenia. marjetka.podobnik@ki.si
Organizational Affiliation: