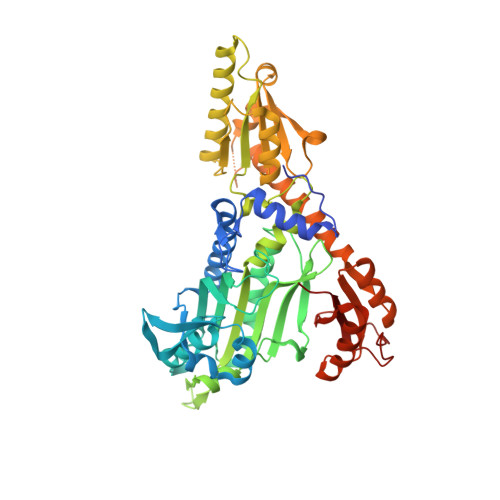
Structure of the prolyl-tRNA synthetase from the eukaryotic pathogen Giardia lamblia.
Larson, E.T., Kim, J.E., Napuli, A.J., Verlinde, C.L., Fan, E., Zucker, F.H., Van Voorhis, W.C., Buckner, F.S., Hol, W.G., Merritt, E.A.(2012) Acta Crystallogr D Biol Crystallogr 68: 1194-1200
- PubMed: 22948920 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444912024699
- Primary Citation Related Structures:
3IAL - PubMed Abstract:
The genome of the human intestinal parasite Giardia lamblia contains only a single aminoacyl-tRNA synthetase gene for each amino acid. The Giardia prolyl-tRNA synthetase gene product was originally misidentified as a dual-specificity Pro/Cys enzyme, in part owing to its unexpectedly high off-target activation of cysteine, but is now believed to be a normal representative of the class of archaeal/eukaryotic prolyl-tRNA synthetases. The 2.2 Å resolution crystal structure of the G. lamblia enzyme presented here is thus the first structure determination of a prolyl-tRNA synthetase from a eukaryote. The relative occupancies of substrate (proline) and product (prolyl-AMP) in the active site are consistent with half-of-the-sites reactivity, as is the observed biphasic thermal denaturation curve for the protein in the presence of proline and MgATP. However, no corresponding induced asymmetry is evident in the structure of the protein. No thermal stabilization is observed in the presence of cysteine and ATP. The implied low affinity for the off-target activation product cysteinyl-AMP suggests that translational fidelity in Giardia is aided by the rapid release of misactivated cysteine.
- Medical Structural Genomics of Pathogenic Protozoa, http://msgpp.org, USA.
Organizational Affiliation: