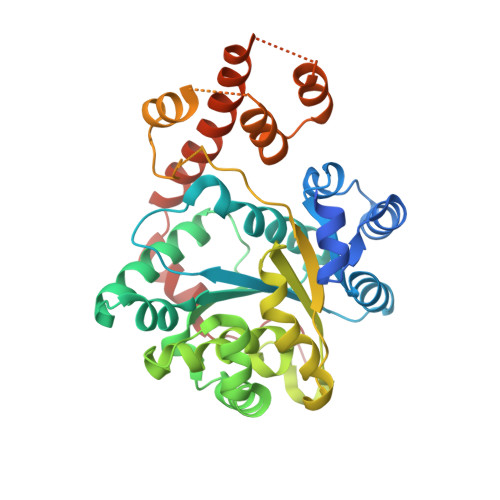
Crystal structures of three protozoan homologs of tryptophanyl-tRNA synthetase.
Merritt, E.A., Arakaki, T.L., Gillespie, R., Napuli, A.J., Kim, J.E., Buckner, F.S., Van Voorhis, W.C., Verlinde, C.L., Fan, E., Zucker, F., Hol, W.G.(2011) Mol Biochem Parasitol 177: 20-28
- PubMed: 21255615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molbiopara.2011.01.003
- Primary Citation Related Structures:
3HV0, 3HZR, 3I05 - PubMed Abstract:
Tryptophanyl-tRNA synthetase (TrpRS) is an essential enzyme that is recognizably conserved across all forms of life. It is responsible for activating and attaching tryptophan to a cognate tRNA(Trp) molecule for use in protein synthesis. In some eukaryotes this original core function has been supplemented or modified through the addition of extra domains or the expression of variant TrpRS isoforms. The three TrpRS structures from pathogenic protozoa described here represent three illustrations of this malleability in eukaryotes. The Cryptosporidium parvum genome contains a single TrpRS gene, which codes for an N-terminal domain of uncertain function in addition to the conserved core TrpRS domains. Sequence analysis indicates that this extra domain, conserved among several apicomplexans, is related to the editing domain of some AlaRS and ThrRS. The C. parvum enzyme remains fully active in charging tRNA(Trp) after truncation of this extra domain. The crystal structure of the active, truncated enzyme is presented here at 2.4Å resolution. The Trypanosoma brucei genome contains separate cytosolic and mitochondrial isoforms of TrpRS that have diverged in their respective tRNA recognition domains. The crystal structure of the T. brucei cytosolic isoform is presented here at 2.8Å resolution. The Entamoeba histolytica genome contains three sequences that appear to be TrpRS homologs. However one of these, whose structure is presented here at 3.0Å resolution, has lost the active site motifs characteristic of the Class I aminoacyl-tRNA synthetase catalytic domain while retaining the conserved features of a fully formed tRNA(Trp) recognition domain. The biological function of this variant E. histolytica TrpRS remains unknown, but, on the basis of a completely conserved tRNA recognition region and evidence for ATP but not tryptophan binding, it is tempting to speculate that it may perform an editing function. Together with a previously reported structure of an unusual TrpRS from Giardia, these protozoan structures broaden our perspective on the extent of structural variation found in eukaryotic TrpRS homologs.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA. merritt@u.washington.edu
Organizational Affiliation: