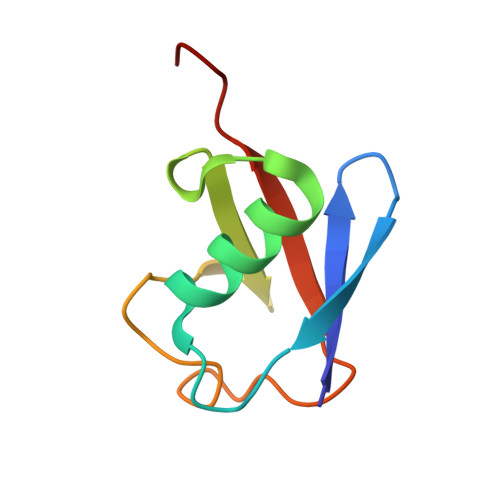
The structure and conformation of Lys63-linked tetraubiquitin.
Datta, A.B., Hura, G.L., Wolberger, C.(2009) J Mol Biology 392: 1117-1124
- PubMed: 19664638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.07.090
- Primary Citation Related Structures:
3HM3 - PubMed Abstract:
Ubiquitination involves the covalent attachment of the ubiquitin (Ub) C-terminus to the lysine side chain of a substrate protein by an isopeptide bond. The modification can comprise a single Ub moiety or a chain of Ub molecules joined by isopeptide bonds between the C-terminus of one Ub with one of the seven lysine residues in the next Ub. Modification of substrate proteins with Lys63-linked poly-Ub plays a key nondegradative signaling role in many biological processes, including DNA repair and nuclear factor-kappaB activation, whereas substrates modified by Lys48-linked chains are targeted to the proteasome for degradation. The distinct signaling properties of alternatively linked Ub chains presumably stem from structural differences that can be distinguished by effector proteins. We have determined the crystal structure of Lys63 tetra-Ub at a resolution of 1.96 A and performed small-angle X-ray scattering experiments and molecular dynamics simulations to probe the conformation of Lys63 tetra-Ub in solution. The chain adopts a highly extended conformation in the crystal, in contrast with the compact globular fold of Lys48 tetra-Ub. Small-angle X-ray scattering experiments show that the Lys63 tetra-Ub chain is dynamic in solution, adopting an ensemble of conformations that are more compact than the extended form in the crystal. The results of these studies provide a basis for understanding the differences in the behavior and recognition of Lys63 poly-Ub chains.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA.
Organizational Affiliation: