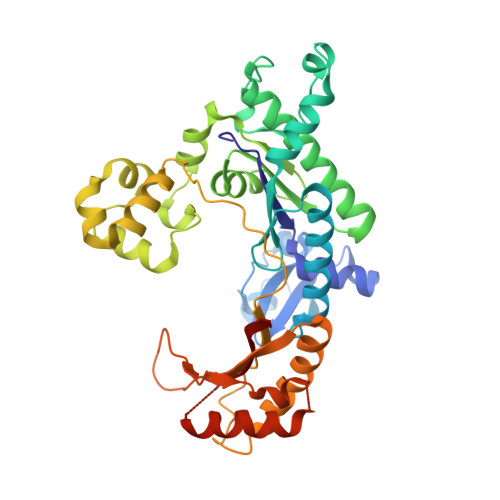
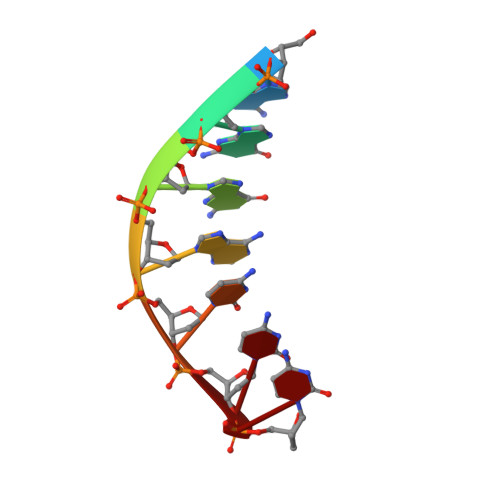
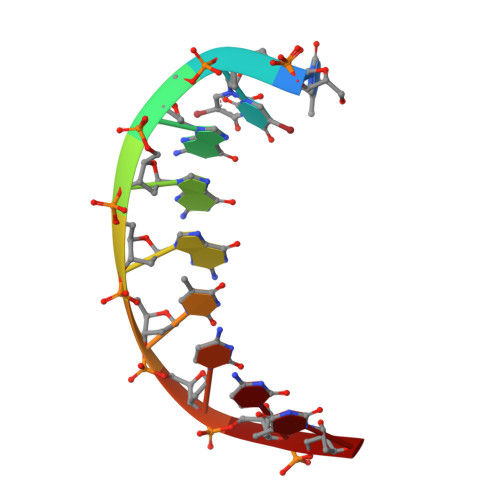
Replication across template T/U by human DNA polymerase-iota.
Jain, R., Nair, D.T., Johnson, R.E., Prakash, L., Prakash, S., Aggarwal, A.K.(2009) Structure 17: 974-980
- PubMed: 19604477 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2009.04.011
- Primary Citation Related Structures:
3H40, 3H4B, 3H4D - PubMed Abstract:
Human DNA polymerase-iota (Poliota) incorporates correct nucleotides opposite template purines with a much higher efficiency and fidelity than opposite template pyrimidines. In fact, the fidelity opposite template T is so poor that Poliota inserts an incorrect dGTP approximately 10 times better than it inserts the correct dATP. We determine here how a template T/U is accommodated in the Poliota active site and why a G is incorporated more efficiently than an A. We show that in the absence of incoming dATP or dGTP (binary complex), template T/U exists in both syn and anti conformations, but in the presence of dATP or dGTP (ternary complexes), template T/U is predominantly in the anti conformation. We also show that dATP and dGTP insert differently opposite template T/U, and that the basis of selection of dGTP over dATP is a hydrogen bond between the N2 amino group of dGTP and Gln59 of Poliota.
- Department of Structural and Chemical Biology, Mount Sinai School of Medicine, New York, NY 10029, USA.
Organizational Affiliation: