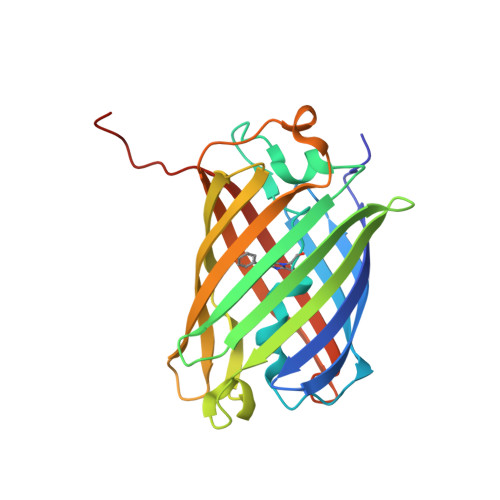
Rotational order-disorder structure of fluorescent protein FP480
Pletnev, S., Morozova, K.S., Verkhusha, V.V., Dauter, Z.(2009) Acta Crystallogr D Biol Crystallogr 65: 906-912
- PubMed: 19690368 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444909020927
- Primary Citation Related Structures:
3H1O, 3H1R - PubMed Abstract:
In the last decade, advances in instrumentation and software development have made crystallography a powerful tool in structural biology. Using this method, structural information can now be acquired from pathological crystals that would have been abandoned in earlier times. In this paper, the order-disorder (OD) structure of fluorescent protein FP480 is discussed. The structure is composed of tetramers with 222 symmetry incorporated into the lattice in two different ways, namely rotated 90 degrees with respect to each other around the crystal c axis, with tetramer axes coincident with crystallographic twofold axes. The random distribution of alternatively oriented tetramers in the crystal creates a rotational OD structure with statistically averaged I422 symmetry, although the presence of very weak and diffuse additional reflections suggests that the randomness is only approximate.
- SAIC-Frederick Inc., Basic Research Program, Argonne National Laboratory, Argonne, IL 60439, USA. svp@ncifcrf.gov
Organizational Affiliation: