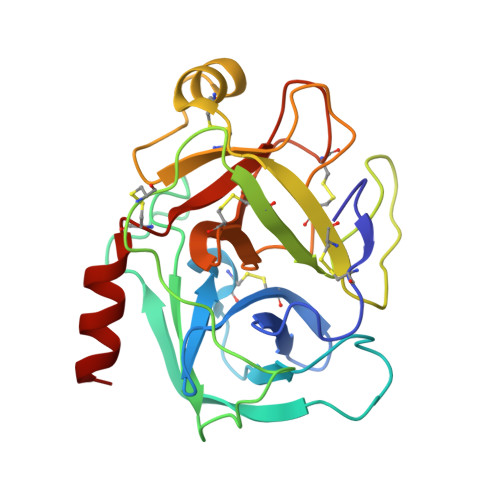
Structural binding evidence of the trypanocidal drugs Berenil and Pentacarinate active principles to a serine protease model.
Perilo, C.S., Pereira, M.T., Santoro, M.M., Nagem, R.A.(2010) Int J Biol Macromol 46: 502-511
- PubMed: 20356563 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2010.03.006
- Primary Citation Related Structures:
3GY2, 3GY3, 3GY4, 3GY5, 3GY6, 3GY7, 3GY8 - PubMed Abstract:
Bovine trypsin is a model system for the serine protease class of enzymes, which is an important target for contemporary medicinal chemistry. Some structural and thermodynamic reports are available on its interaction with benzamidine-based compounds but no structural information is available so far on its binding modes to the active principles of the trypanocidal drugs Pentacarinate (pentamidine) and Berenil (diminazene). The crystallographic structures of bovine beta-trypsin in complex with the ligands were determined to a resolution of 1.57 A (diminazene) and 1.70 A (diminazene and pentamidine). The second benzamidine moieties in these inhibitors are bound to the enzyme in different hot spots and only few hydrogen bonds mediate these interactions. Thermodynamic parameters for the association of pentamidine with beta-trypsin reveal that this inhibitor has about 1.3-fold lower affinity than diminazene. Moreover its binding mode resembles other benzamidine-based compounds that assess the aryl binding pocket of the enzyme; however, with almost 2.5-fold higher affinity. This is the first structural evidence of the binding of Berenil and Pentacarinate active principles trypanocidal drugs to serine proteases.
- Departamento de Bioquímica e Imunologia, Instituto de Ciências Biológicas, Universidade Federal de Minas Gerais, Avenida Antônio Carlos 6627, Belo Horizonte, MG, CEP 31270-901, Brazil.
Organizational Affiliation: