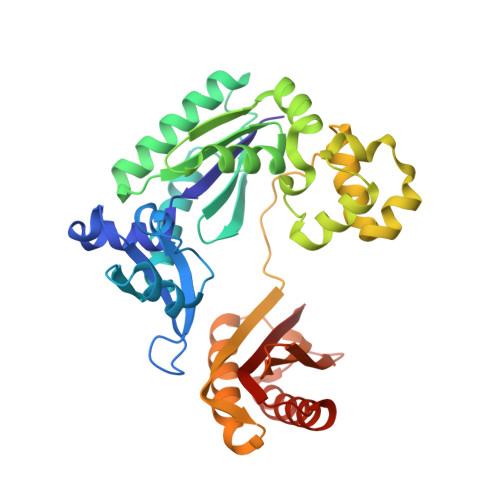
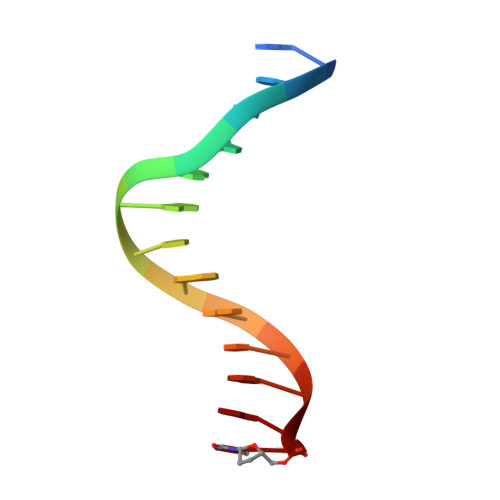
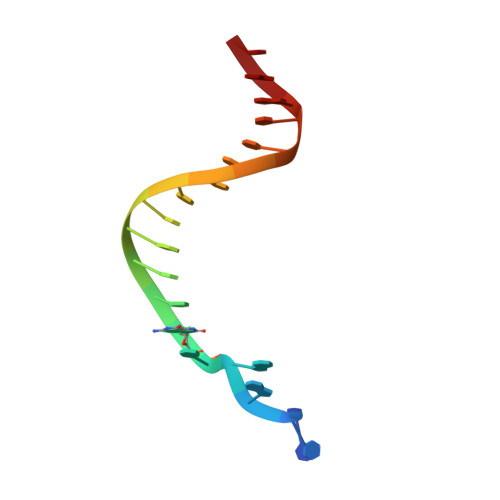
Impact of conformational heterogeneity of OxoG lesions and their pairing partners on bypass fidelity by Y family polymerases.
Rechkoblit, O., Malinina, L., Cheng, Y., Geacintov, N.E., Broyde, S., Patel, D.J.(2009) Structure 17: 725-736
- PubMed: 19446528 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2009.03.011
- Primary Citation Related Structures:
3GII, 3GIJ, 3GIK, 3GIL, 3GIM - PubMed Abstract:
7,8-Dihydro-8-oxoguanine (oxoG), the predominant oxidative DNA damage lesion, is processed differently by high-fidelity and Y-family lesion bypass polymerases. Although high-fidelity polymerases extend predominantly from an A base opposite an oxoG, the Y-family polymerases Dpo4 and human Pol eta preferentially extend from the oxoG*C base pair. We have determined crystal structures of extension Dpo4 ternary complexes with oxoG opposite C, A, G, or T and the next nascent base pair. We demonstrate that neither template backbone nor the architecture of the active site is perturbed by the oxoG(anti)*C and oxoG*A pairs. However, the latter manifest conformational heterogeneity, adopting both oxoG(syn)*A(anti) and oxoG(anti)*A(syn) alignment. Hence, the observed reduced primer extension from the dynamically flexible 3'-terminal primer base A is explained. Because of homology between Dpo4 and Pol eta, such a dynamic screening mechanism might be utilized by Dpo4 and Pol eta to regulate error-free versus error-prone bypass of oxoG and other lesions.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10021, USA.
Organizational Affiliation: