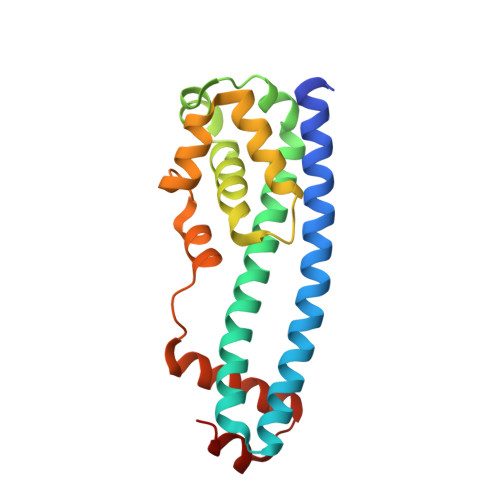
Structural basis of Ist1 function and Ist1-Did2 interaction in the multivesicular body pathway and cytokinesis.
Xiao, J., Chen, X.W., Davies, B.A., Saltiel, A.R., Katzmann, D.J., Xu, Z.(2009) Mol Biol Cell 20: 3514-3524
- PubMed: 19477918 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1091/mbc.e09-05-0403
- Primary Citation Related Structures:
3GGY, 3GGZ - PubMed Abstract:
The ESCRT machinery functions in several important eukaryotic cellular processes. The AAA-ATPase Vps4 catalyzes disassembly of the ESCRT-III complex and may regulate membrane deformation and vesicle scission as well. Ist1 was proposed to be a regulator of Vps4, but its mechanism of action was unclear. The crystal structure of the N-terminal domain of Ist1 (Ist1NTD) reveals an ESCRT-III subunit-like fold, implicating Ist1 as a divergent ESCRT-III family member. Ist1NTD specifically binds to the ESCRT-III subunit Did2, and cocrystallization of Ist1NTD with a Did2 fragment shows that Ist1 interacts with the Did2 C-terminal MIM1 (MIT-interacting motif 1) via a novel MIM-binding structural motif. This arrangement indicates a mechanism for intermolecular ESCRT-III subunit association and may also suggest one form of ESCRT-III subunit autoinhibition via intramolecular interaction.
- Life Sciences Institute and Department of Biological Chemistry, Department of Molecular and Integrative Physiology, Medical School, University of Michigan, Ann Arbor, MI 48109, USA.
Organizational Affiliation: