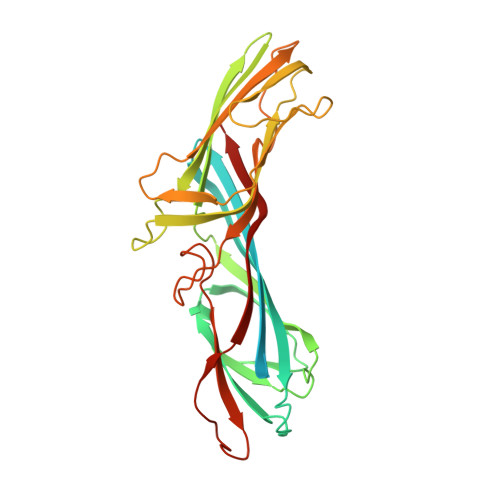
Syp1 is a conserved endocytic adaptor that contains domains involved in cargo selection and membrane tubulation.
Reider, A., Barker, S.L., Mishra, S.K., Im, Y.J., Maldonado-Baez, L., Hurley, J.H., Traub, L.M., Wendland, B.(2009) EMBO J 28: 3103-3116
- PubMed: 19713939 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2009.248
- Primary Citation Related Structures:
3G9G, 3G9H - PubMed Abstract:
Internalization of diverse transmembrane cargos from the plasma membrane requires a similarly diverse array of specialized adaptors, yet only a few adaptors have been characterized. We report the identification of the muniscin family of endocytic adaptors that is conserved from yeast to human beings. Solving the structures of yeast muniscin domains confirmed the unique combination of an N-terminal domain homologous to the crescent-shaped membrane-tubulating EFC/F-BAR domains and a C-terminal domain homologous to cargo-binding mu homology domains (muHDs). In vitro and in vivo assays confirmed membrane-tubulation activity for muniscin EFC/F-BAR domains. The muHD domain has conserved interactions with the endocytic adaptor/scaffold Ede1/eps15, which influences muniscin localization. The transmembrane protein Mid2, earlier implicated in polarized Rho1 signalling, was identified as a cargo of the yeast adaptor protein. These and other data suggest a model in which the muniscins provide a combined adaptor/membrane-tubulation activity that is important for regulating endocytosis.
- Department of Biology, The Johns Hopkins University, Baltimore, MD 21218-2685, USA.
Organizational Affiliation: