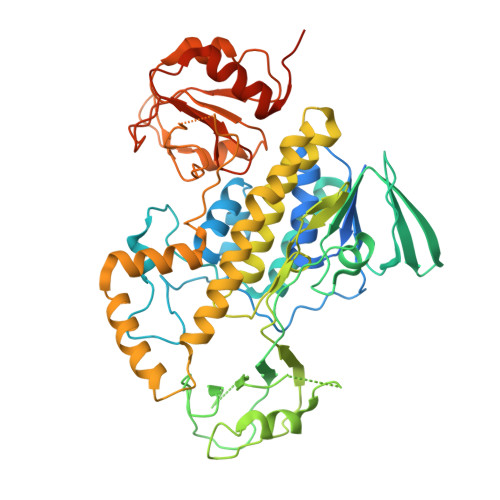
Crystal structure of Baeyer-Villiger monooxygenase MtmOIV, the key enzyme of the mithramycin biosynthetic pathway .
Beam, M.P., Bosserman, M.A., Noinaj, N., Wehenkel, M., Rohr, J.(2009) Biochemistry 48: 4476-4487
- PubMed: 19364090 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi8023509
- Primary Citation Related Structures:
3FMW - PubMed Abstract:
Baeyer-Villiger monooxygenases (BVMOs), mostly flavoproteins, were shown to be powerful biocatalysts for synthetic organic chemistry applications and were also suggested to play key roles for the biosyntheses of various natural products. Here we present the three-dimensional structure of MtmOIV, a 56 kDa homodimeric FAD- and NADPH-dependent monooxygenase, which catalyzes the key frame-modifying step of the mithramycin biosynthetic pathway and currently the only BVMO proven to react with its natural substrate via a Baeyer-Villiger reaction. MtmOIV's structure was determined by X-ray crystallography using molecular replacement to a resolution of 2.9 A. MtmOIV cleaves a C-C bond, essential for the conversion of the biologically inactive precursor, premithramycin B, into the active drug mithramycin. The MtmOIV structure combined with substrate docking calculations and site-directed mutagenesis experiments identifies several residues that participate in cofactor and substrate binding. Future experimentation aimed at broadening the substrate specificity of the enzyme could facilitate the generation of chemically diverse mithramycin analogues through combinatorial biosynthesis.
- Department of Pharmaceutical Sciences, College of Pharmacy, and Kentucky Center for Structural Biology, University of Kentucky, Lexington, Kentucky 40536, USA.
Organizational Affiliation: