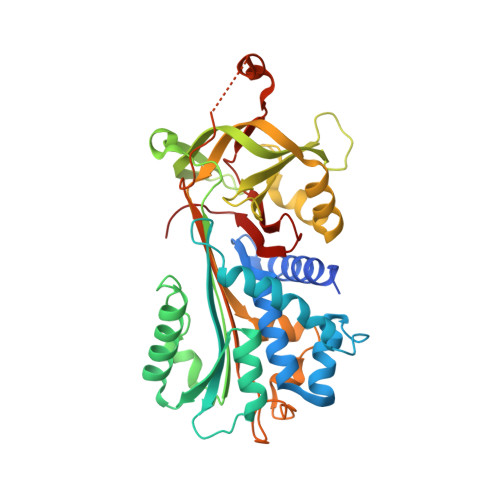
The 2.1-A crystal structure of native neuroserpin reveals unique structural elements that contribute to conformational instability
Takehara, S., Onda, M., Zhang, J., Nishiyama, M., Yang, X., Mikami, B., Lomas, D.A.(2009) J Mol Biology 388: 11-20
- PubMed: 19285087 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.03.007
- Primary Citation Related Structures:
3FGQ - PubMed Abstract:
Neuroserpin is a selective inhibitor of tissue-type plasminogen activator (tPA) that plays an important role in neuronal plasticity, memory, and learning. We report here the crystal structure of native human neuroserpin at 2.1 A resolution. The structure has a helical reactive center loop and an omega loop between strands 1B and 2B. The omega loop contributes to the inhibition of tPA, as deletion of this motif reduced the association rate constant with tPA by threefold but had no effect on the kinetics of interaction with urokinase. Point mutations in neuroserpin cause the formation of ordered intracellular polymers that underlie dementia familial encephalopathy with neuroserpin inclusion bodies (FENIB). Wild-type neuroserpin is also unstable and readily forms polymers under near-physiological conditions in vitro. This is, in part, due to the substitution of a conserved alanine for serine at position 340. The replacement of Ser340 by Ala increased the melting temperature by 3 degrees C and reduced polymerization as compared to wild-type neuroserpin. Similarly, neuroserpin has Asn-Leu-Val at the end of helix F and thus differs markedly from the Gly-X-Ile consensus sequence of the serpins. Restoration of these amino acids to the consensus sequence increased thermal stability and reduced the polymerization of neuroserpin and its transition to the latent conformer. Moreover, introduction of the consensus sequence into S49P neuroserpin that causes FENIB increased the stability and inhibitory activity of the mutant, as well as blocked polymerization and increased the yield of protein during refolding. These data provide a molecular explanation for the inherent instability of neuroserpin and the effect of point mutations that underlie the dementia FENIB.
- Division of Applied Life Sciences, The Graduate School of Agriculture, Kyoto University, Uji 611-0011, Japan.
Organizational Affiliation: