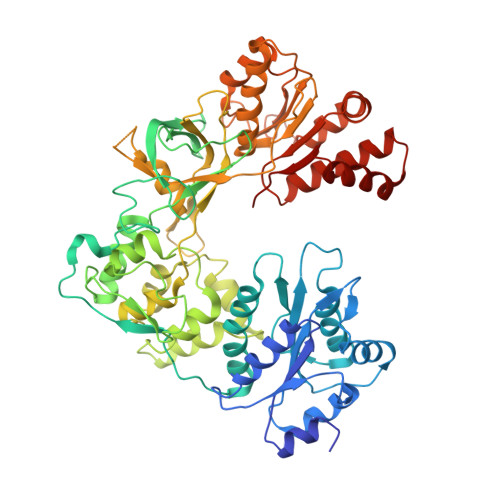
Structure and function of an NADPH-cytochrome P450 oxidoreductase in an open conformation capable of reducing cytochrome P450
Hamdane, D., Xia, C., Im, S.C., Zhang, H., Kim, J.J., Waskell, L.(2009) J Biological Chem 284: 11374-11384
- PubMed: 19171935 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M807868200
- Primary Citation Related Structures:
3ES9 - PubMed Abstract:
NADPH-cytochrome P450 oxidoreductase (CYPOR) catalyzes the transfer of electrons to all known microsomal cytochromes P450. A CYPOR variant, with a 4-amino acid deletion in the hinge connecting the FMN domain to the rest of the protein, has been crystallized in three remarkably extended conformations. The variant donates an electron to cytochrome P450 at the same rate as the wild-type, when provided with sufficient electrons. Nevertheless, it is defective in its ability to transfer electrons intramolecularly from FAD to FMN. The three extended CYPOR structures demonstrate that, by pivoting on the C terminus of the hinge, the FMN domain of the enzyme undergoes a structural rearrangement that separates it from FAD and exposes the FMN, allowing it to interact with its redox partners. A similar movement most likely occurs in the wild-type enzyme in the course of transferring electrons from FAD to its physiological partner, cytochrome P450. A model of the complex between an open conformation of CYPOR and cytochrome P450 is presented that satisfies mutagenesis constraints. Neither lengthening the linker nor mutating its sequence influenced the activity of CYPOR. It is likely that the analogous linker in other members of the diflavin family functions in a similar manner.
- University of Michigan Medical School and Veterans Affairs Medical Research Center, Ann Arbor, Michigan 48105, USA.
Organizational Affiliation: