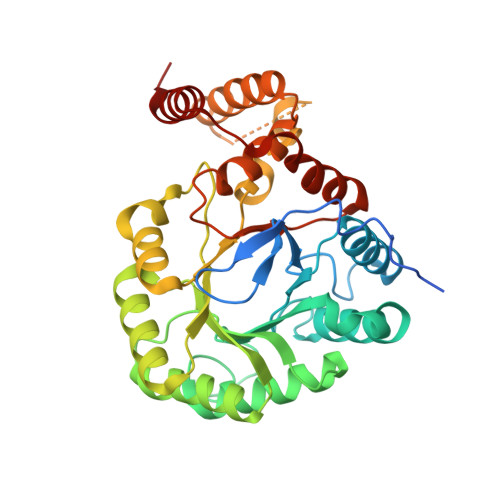
Structural and biochemical studies of novel aldo-keto reductases for the biocatalytic conversion of 3-hydroxybutanal to 1,3-butanediol.
Kim, T., Flick, R., Brunzelle, J., Singer, A., Evdokimova, E., Brown, G., Joo, J.C., Minasov, G.A., Anderson, W.F., Mahadevan, R., Savchenko, A., Yakunin, A.F.(2017) Appl Environ Microbiol
- PubMed: 28130301 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AEM.03172-16
- Primary Citation Related Structures:
3ERP, 5T79 - PubMed Abstract:
The nonnatural alcohol 1,3-butanediol (1,3-BDO) is a valuable building block for the synthesis of various polymers. One of the potential pathways for the biosynthesis of 1,3-BDO includes the biotransformation of acetaldehyde to 1,3-BDO via 3-hydroxybutanal (3-HB) using aldolases and aldo-keto reductases (AKRs). This pathway requires an AKR selective for 3-HB, but inactive toward acetaldehyde, so it can be used for one-pot synthesis. In this work, we screened more than 20 purified uncharacterized AKRs for 3-HB reduction and identified 10 enzymes with significant activity and nine proteins with detectable activity. PA1127 from Pseudomonas aeruginosa showed the highest activity and was selected for comparative studies with STM2406 from Salmonella enterica serovar Typhimurium, for which we have determined the crystal structure. Both AKRs used NADPH as a cofactor, reduced a broad range of aldehydes, and showed low activities toward acetaldehyde. The crystal structures of STM2406 in complex with cacodylate or NADPH revealed the active site with bound molecules of a substrate mimic or cofactor. Site-directed mutagenesis of STM2406 and PA1127 identified the key residues important for the activity against 3-HB and aromatic aldehydes, which include the residues of the substrate-binding pocket and C-terminal loop. Our results revealed that the replacement of the STM2406 Asn65 by Met enhanced the activity and the affinity of this protein toward 3-HB, resulting in a 7-fold increase in k cat / K m Our work provides further insights into the molecular mechanisms of the substrate selectivity of AKRs and for the rational design of these enzymes toward new substrates. IMPORTANCE In this study, we identified several aldo-keto reductases with significant activity in reducing 3-hydroxybutanal to 1,3-butanediol (1,3-BDO), an important commodity chemical. Biochemical and structural studies of these enzymes revealed the key catalytic and substrate-binding residues, including the two structural determinants necessary for high activity in the biosynthesis of 1,3-BDO. This work expands our understanding of the molecular mechanisms of the substrate selectivity of aldo-keto reductases and demonstrates the potential for protein engineering of these enzymes for applications in the biocatalytic production of 1,3-BDO and other valuable chemicals.
- Department of Chemical Engineering and Applied Chemistry, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: