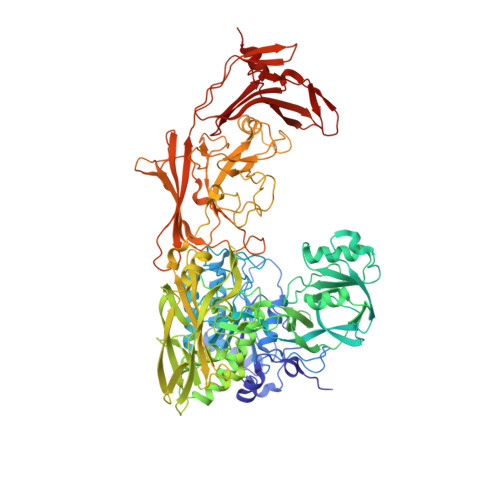
Model for Substrate Interactions in C5a Peptidase from Streptococcus pyogenes: A 1.9 A Crystal Structure of the Active Form of ScpA
Kagawa, T.F., O'Connell, M.R., Mouat, P., Paoli, M., O'Toole, P.W., Cooney, J.C.(2009) J Mol Biology 386: 754-772
- PubMed: 19152799 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.12.074
- Primary Citation Related Structures:
3EIF - PubMed Abstract:
The crystal structure of an active form of ScpA has been solved to 1.9 A resolution. ScpA is a multidomain cell-envelope subtilase from Streptococcus pyogenes that cleaves complement component C5a. The catalytic triad of ScpA is geometrically consistent with other subtilases, clearly demonstrating that the additional activation mechanism proposed for the Streptococcus agalactiae homologue (ScpB) is not required for ScpA. The ScpA structure revealed that access to the catalytic site is restricted by variable regions in the catalytic domain (vr7, vr9, and vr11) and by the presence of the inserted protease-associated (PA) domain and the second fibronectin type III domains (Fn2). Modeling of the ScpA-C5a complex indicates that the substrate binds with carboxyl-terminal residues (65-74) extended through the active site and core residues (1-64) forming exosite-type interactions with the Fn2 domain. This is reminiscent of the two-site mechanism proposed for C5a binding to its receptor. In the nonprime region of the active site, interactions with the substrate backbone are predicted to be more similar to those observed in kexins, involving a single beta-strand in the peptidase. However, in contrast to kexins, there would be diminished emphasis on side-chain interactions, with little charged character in the S3-S1 and S6-S4 subsites occupied by the side chains of residues in vr7 and vr9. Substrate binding is anticipated to be dominated by ionic interactions in two distinct regions of ScpA. On the prime side of the active site, salt bridges are predicted between P1', P2', and P7' residues, and residues in the catalytic and PA domains. Remote to the active site, a larger number of ionic interactions between residues in the C5a core and the Fn2 domain are observed in the model. Thus, both PA and Fn2 domains are expected to play significant roles in substrate recognition.
- Department of Life Sciences, University of Limerick, Limerick, Ireland. kagawatf@gmail.com
Organizational Affiliation: