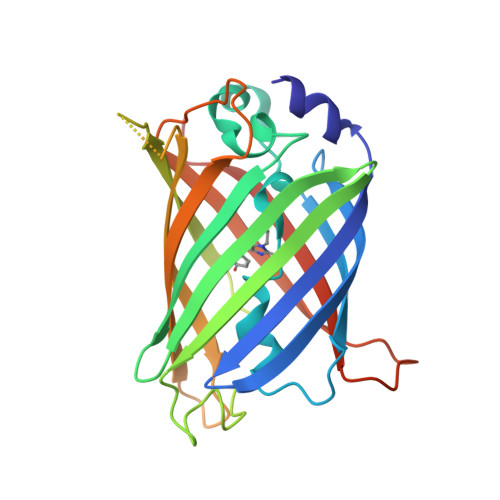
Applicability of superfolder YFP bimolecular fluorescence complementation in vitro.
Ottmann, C., Weyand, M., Wolf, A., Kuhlmann, J., Ottmann, C.(2009) Biol Chem 390: 81-90
- PubMed: 19007309 Search on PubMed
- DOI: https://doi.org/10.1515/BC.2009.008
- Primary Citation Related Structures:
3ED8 - PubMed Abstract:
Bimolecular fluorescence complementation (BiFC) using yellow fluorescent protein (YFP) is a widely employed method to study protein-protein interactions in cells. As yet, this technique has not been used in vitro. To evaluate a possible application of BiFC in vitro, we constructed a 'superfolder split YFP' system where 15 mutations enhance expression of the fusion proteins in Escherichia coli and enable a native purification due to improved solubility. Here, we present the crystal structure of 'superfolder YFP', providing the structural basis for the enhanced folding and stability characteristics. Complementation between the two non-fluorescent YFP fragments fused to HRas and Raf1RBD or to 14-3-3 and PMA2-CT52 resulted in the constitution of the functional fluorophore. The in vivo BiFC with these protein interaction pairs was demonstrated in eukaryotic cell lines as well. Here, we present for the first time BiFC in vitro studies with natively purified superfolder YFP fusion proteins and show the potential and drawbacks of this method for analyzing protein-protein interactions.
- Department of Structural Biology, Max Planck Institute of Molecular Physiology, Otto-Hahn-Str. 11, D-44227 Dortmund, Germany.
Organizational Affiliation: