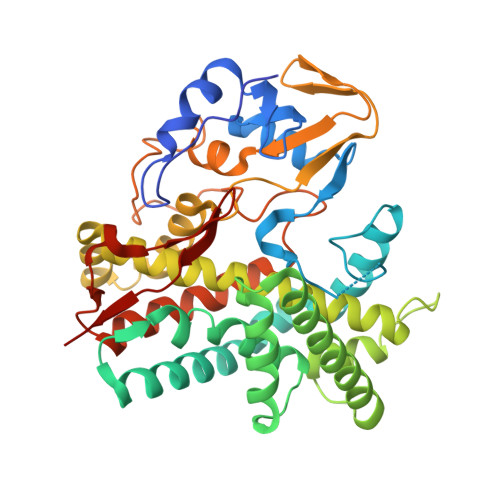
Crystal structures of cytochrome P450 105P1 from Streptomyces avermitilis: conformational flexibility and histidine ligation state
Xu, L.H., Fushinobu, S., Ikeda, H., Wakagi, T., Shoun, H.(2009) J Bacteriol 191: 1211-1219
- PubMed: 19074393 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.01276-08
- Primary Citation Related Structures:
3E5J, 3E5K, 3E5L - PubMed Abstract:
The polyene macrolide antibiotic filipin is widely used as a probe for cholesterol in biological membranes. The filipin biosynthetic pathway of Streptomyces avermitilis contains two position-specific hydroxylases, C26-specific CYP105P1 and C1'-specific CYP105D6. In this study, we describe the three X-ray crystal structures of CYP105P1: the ligand-free wild-type (WT-free), 4-phenylimidazole-bound wild-type (WT-4PI), and ligand-free H72A mutant (H72A-free) forms. The BC loop region in the WT-free structure has a unique feature; the side chain of His72 within this region is ligated to the heme iron. On the other hand, this region is highly disordered and widely open in WT-4PI and H72A-free structures, respectively. Histidine ligation of wild-type CYP105P1 was not detectable in solution, and a type II spectral change was clearly observed when 4-phenylimidazole was titrated. The H72A mutant showed spectroscopic characteristics that were almost identical to those of the wild-type protein. In the H72A-free structure, there is a large pocket that is of the same size as the filipin molecule. The highly flexible feature of the BC loop region of CYP105P1 may be required to accept a large hydrophobic substrate.
- Department of Biotechnology, Graduate School of Agriculture and Life Sciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: