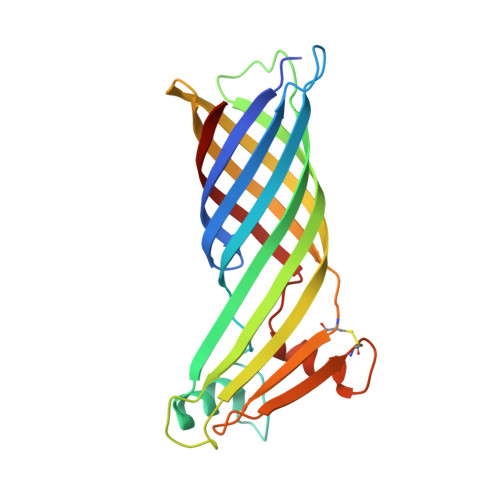
Crystal structure of a major outer membrane protein from Thermus thermophilus HB27
Brosig, A., Nesper, J., Boos, W., Welte, W., Diederichs, K.(2009) J Mol Biology 385: 1445-1455
- PubMed: 19101566 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.12.003
- Primary Citation Related Structures:
3DZM - PubMed Abstract:
The thermophilic eubacterium Thermus thermophilus belongs to one of the oldest branches of evolution and has a multilayered cell envelope that differs from that of modern Gram-negative bacteria. Its outer membrane contains integral outer membrane proteins (OMPs), of which only a few are characterized. TtoA, a new beta-barrel OMP, was identified by searching the genome sequence of strain HB27 for the presence of a C-terminal signature sequence. The structure of TtoA was determined to a resolution of 2.8 A, representing the first crystal structure of an OMP from a thermophilic bacterium. TtoA consists of an eight-stranded beta-barrel with a large extracellular part to which a divalent cation is bound. A five-stranded extracellular beta-sheet protrudes out of the membrane-embedded transmembrane barrel and is stabilized by a disulfide bridge. The edge of this beta-sheet forms crystal contacts that could mimic interactions with other proteins. In modern Gram-negative bacteria, the C-terminal signature sequence of OMPs is required for binding to an Omp85 family protein as a prerequisite for its assembly. We present hints that a similar assembly pathway exists in T. thermophilus by an in vitro binding assay, where unfolded TtoA binds to the Thermus Omp85 family protein TtOmp85, while a mutant without the signature sequence does not.
- Department of Biology, University of Konstanz, Konstanz, Germany.
Organizational Affiliation: