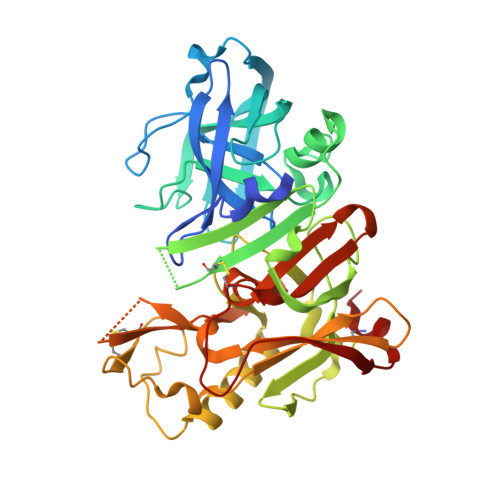
Structure-based design and synthesis of macrocyclic peptidomimetic beta-secretase (BACE-1) inhibitors.
Machauer, R., Veenstra, S., Rondeau, J.M., Tintelnot-Blomley, M., Betschart, C., Neumann, U., Paganetti, P.(2009) Bioorg Med Chem Lett 19: 1361-1365
- PubMed: 19195886 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2009.01.036
- Primary Citation Related Structures:
3DUY - PubMed Abstract:
The hydroxyethylene octapeptide inhibitor OM99-2 served as starting point to create the tripeptide inhibitor 1 and its analogues 2a and b. An X-ray co-crystal structure of 1 with BACE-1 allowed the design and syntheses of a series of macrocyclic analogues 3a-h covalently linking the P1 and P3 side-chains. These inhibitors show improved enzymatic potency over their open-chain analogue. Inhibitor 3h also shows activity in a cellular system.
- Novartis Institutes for BioMedical Research, Novartis Pharma AG, PO Box, CH-4002 Basel, Switzerland.
Organizational Affiliation: