Structural details of HIV-1 recognition by the broadly neutralizing monoclonal antibody 2F5: epitope conformation, antigen-recognition loop mobility, and anion-binding site.
Julien, J.P., Bryson, S., Nieva, J.L., Pai, E.F.(2008) J Mol Biology 384: 377-392
- PubMed: 18824005 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.09.024
- Primary Citation Related Structures:
2P8L, 2P8M, 2P8P, 2PR4, 3D0L, 3D0V, 3DRO, 3DRQ - PubMed Abstract:
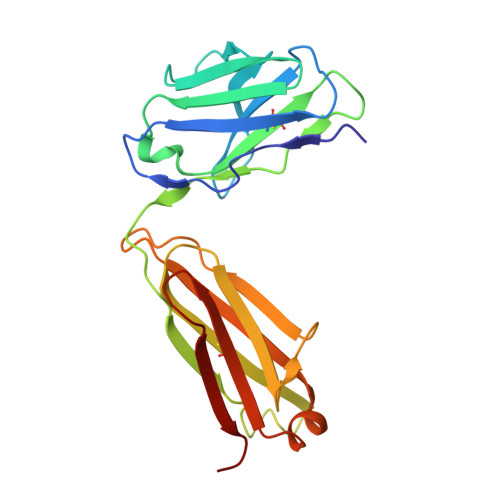
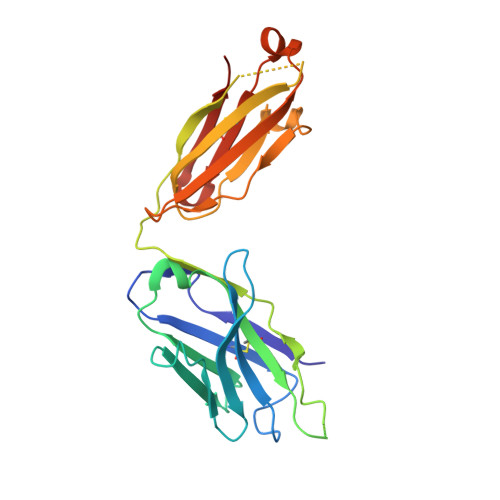
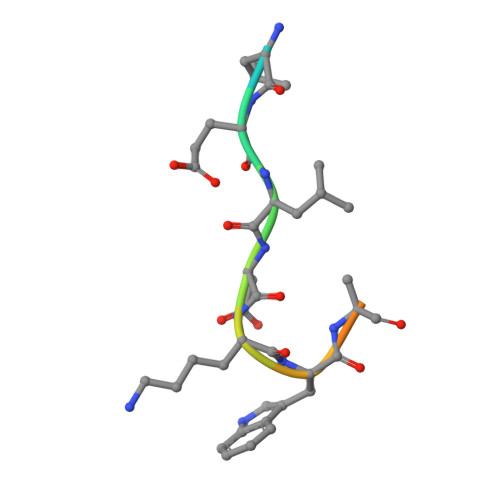
2F5 is a monoclonal antibody with potent and broadly neutralizing activity against HIV-1. It targets the membrane-proximal external region (MPER) of the gp41 subunit of the envelope glycoprotein and interferes with the process of fusion between viral and host cell membranes. This study presents eight 2F5 F(ab)' crystal structures in complex with various gp41 peptide epitopes. These structures reveal several key features of this antibody-antigen interaction. (1) Whenever free of contacts caused by crystal artifacts, the extended complementarity-determining region H3 loop is mobile; this is true for ligand-free and epitope-bound forms. (2) The interaction between the antibody and the gp41 ELDKWA epitope core is absolutely critical, and there are also close and specific contacts with residues located N-terminal to the epitope core. (3) Residues located at the C-terminus of the gp41 ELDKWA core do not interact as tightly with the antibody. However, in the presence of a larger peptide containing the gp41 fusion peptide segment, these residues adopt a conformation consistent with the start of an alpha-helix. (4) At high sulfate concentrations, the electron density maps of 2F5 F(ab)'-peptide complexes contain a peak that may mark a binding site for phosphate groups of negatively charged lipid headgroups. The refined atomic-level details of 2F5 paratope-epitope interactions revealed here should contribute to a better understanding of the mechanism of 2F5-based virus neutralization, in general, and prove important for the design of potential vaccine candidates intended to elicit 2F5-like antibody production.
- Department of Biochemistry, University of Toronto, 1 King's College Circle, Toronto, Ontario, Canada M5S 1A8.
Organizational Affiliation: